| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,706,492 – 18,706,582 |
| Length | 90 |
| Max. P | 0.979770 |

| Location | 18,706,492 – 18,706,582 |
|---|---|
| Length | 90 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 62.56 |
| Shannon entropy | 0.65898 |
| G+C content | 0.48839 |
| Mean single sequence MFE | -26.44 |
| Consensus MFE | -12.10 |
| Energy contribution | -13.70 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.979770 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
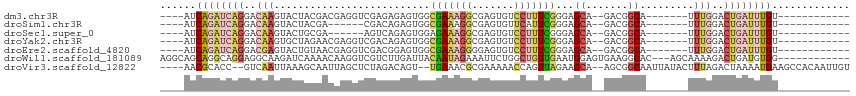
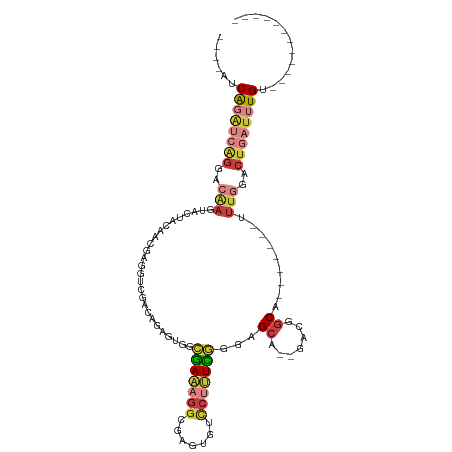
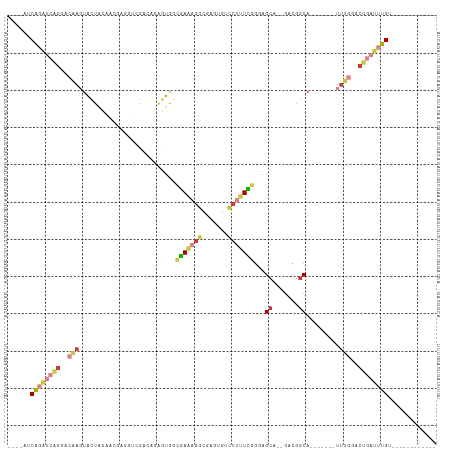
>dm3.chr3R 18706492 90 - 27905053 ----AUCAGAUCAGGACAAGUACUACGACGAGGUCGAGAGAGUGGCGAAAGGCGAGUGUCCUUUCGGGAGCA--GACGGCA-------UUUGGACUGAUUUGU------------ ----..((((((((..(((((.((.((((...)))).....((..((((((((....).)))))))...)).--...)).)-------))))..)))))))).------------ ( -29.40, z-score = -1.97, R) >droSim1.chr3R 18521524 84 - 27517382 ----AUCAGAUCAGGACAAGUACUACGA------CGACAGAGUGGCGAAAGGCGAGUGUUCAUUCGGGAGCA--GACGGCA-------UUUGGACUGAUUUGU------------ ----..((((((((..(((((.((((..------.......)))).....(.((..(((((......)))))--..)).))-------))))..)))))))).------------ ( -26.50, z-score = -2.75, R) >droSec1.super_0 19075420 84 - 21120651 ----AUCAGAUCAGGACAAGUACUGCGA------AGUCAGAGUGGAGAAAGGCGAGUGUCCUUUCGGGAGCA--GACGGCA-------UUUGGACUGAUUUGU------------ ----..((((((((..((((...(((..------.(((...((...(((((((....).))))))....)).--))).)))-------))))..)))))))).------------ ( -26.20, z-score = -1.75, R) >droYak2.chr3R 19563810 90 - 28832112 ----AUCAGAUCAGGACAAGUGCUAGAACGAGGUCGACAGAGUGGCGAAAGGCGAGUGUCCUUUCGGGAGCA--GACGGCA-------UUUGGACUGAUUUGU------------ ----..((((((((..((((((((....(((((..((((...((.(....).))..)))).)))))(.....--..)))))-------))))..)))))))).------------ ( -33.90, z-score = -3.39, R) >droEre2.scaffold_4820 1110604 90 + 10470090 ----AUCAGAUCAGGACGAGUACUGUAACGAGGUCGACGGAGUGGCGAAAGGGGAGUGUCCUUUCGGGAGCA--GACGGCA-------UUUGGACUGAUUUGU------------ ----..((((((((..(((((.((((..((....)).....((..((((((((.....))))))))...)).--.)))).)-------))))..)))))))).------------ ( -31.50, z-score = -2.57, R) >droWil1.scaffold_181089 11045493 100 + 12369635 AGGCAGCAGGCAGGAGGCAAGAUCAAAACAAGGUCGUCUUGAUUACAAUAGAAAUUCUGGCUGUUGAAUGGAGUGAAGGCAC---AGCAAAAGACUGAUGUGG------------ ...((((((.(((((.....((((((.((......)).))))))..........))))).)))))).............(((---(.((......)).)))).------------ ( -22.46, z-score = -0.15, R) >droVir3.scaffold_12822 3923444 105 + 4096053 ----AACGCACC--GUCAAUUAAAGCAAUUAGCUCUAGACAGU--UGAAACGCGAAAAACCAGUUAGAAGCA--AGCGGCAAUUAUACUUUAGACUAAAAUGAAGCCACAAUUGU ----..(((...--((........)).....(((((((...((--(...........)))...)))).))).--.)))((((((...(((((........)))))....)))))) ( -15.10, z-score = 0.51, R) >consensus ____AUCAGAUCAGGACAAGUACUACAACGAGGUCGACAGAGUGGCGAAAGGCGAGUGUCCUUUCGGGAGCA__GACGGCA_______UUUGGACUGAUUUGU____________ ......((((((((..(((..........................(((((((.......)))))))...((.......)).........)))..))))))))............. (-12.10 = -13.70 + 1.60)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:36:22 2011