
| Sequence ID | dm3.chr2L |
|---|---|
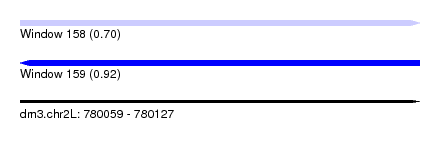
| Location | 780,059 – 780,127 |
| Length | 68 |
| Max. P | 0.923779 |

| Location | 780,059 – 780,127 |
|---|---|
| Length | 68 |
| Sequences | 5 |
| Columns | 68 |
| Reading direction | forward |
| Mean pairwise identity | 73.09 |
| Shannon entropy | 0.48881 |
| G+C content | 0.54832 |
| Mean single sequence MFE | -19.64 |
| Consensus MFE | -12.32 |
| Energy contribution | -12.68 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.695251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
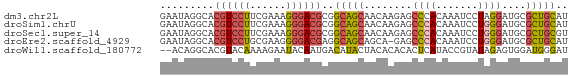
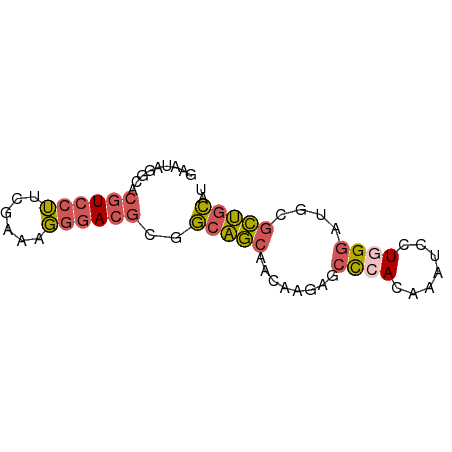
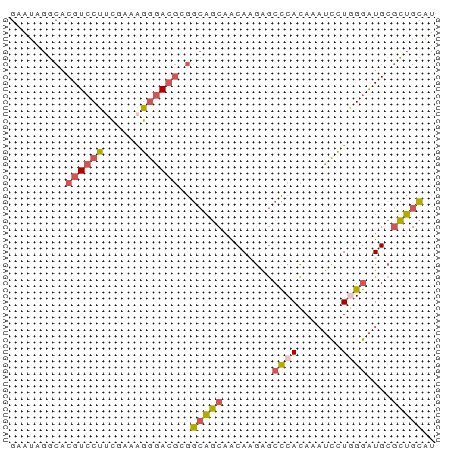
>dm3.chr2L 780059 68 + 23011544 GAAUAGGCACGUCCUUCGAAAGGGACGCGGCAGCAACAAGAGCCCACAAAUCCUAGGAUGCGCUGCAU .......(.((((((......)))))).)(((((.....(....)....(((....)))..))))).. ( -18.70, z-score = -0.63, R) >droSim1.chrU 11632596 68 + 15797150 GAAUAGGCACGUCCUUCGAAAGGGACGCGGCAGCAACAAGAGCCCACAAAUCCUGGGAUGCGCUGCAU .......(.((((((......)))))).)(((((..((....((((.......)))).)).))))).. ( -22.10, z-score = -1.35, R) >droSec1.super_14 750668 68 + 2068291 GAAUAGGCACGUCCUUCGAAAGGGACGCGGCAGCAACAAGAGCCCACAAAUCCUGGGAUGCGCUGCGU .......(.((((((......)))))).)(((((..((....((((.......)))).)).))))).. ( -22.10, z-score = -0.92, R) >droEre2.scaffold_4929 832790 67 + 26641161 GAAUAGGCACGUCCUGCGAAGGGGACGAGGCAGCAGCA-GAGCCCACAAAUCCUGGGAUGCGCUGCAU .........((((((......))))))..(((((.(((-...((((.......)))).)))))))).. ( -27.50, z-score = -2.41, R) >droWil1.scaffold_180772 6452536 66 - 8906247 --ACAGGCACGUACAAAAGAAUACAAUGACAUACUACACACACUCAUACCGUAUAGAGUGGAUGGGAU --..................................((..(((((..........)))))..)).... ( -7.80, z-score = 0.16, R) >consensus GAAUAGGCACGUCCUUCGAAAGGGACGCGGCAGCAACAAGAGCCCACAAAUCCUGGGAUGCGCUGCAU .........((((((......))))))..(((((........((((.......))))....))))).. (-12.32 = -12.68 + 0.36)
| Location | 780,059 – 780,127 |
|---|---|
| Length | 68 |
| Sequences | 5 |
| Columns | 68 |
| Reading direction | reverse |
| Mean pairwise identity | 73.09 |
| Shannon entropy | 0.48881 |
| G+C content | 0.54832 |
| Mean single sequence MFE | -21.02 |
| Consensus MFE | -15.94 |
| Energy contribution | -15.58 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.923779 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
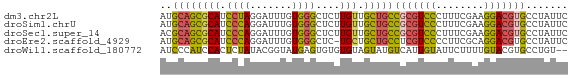
>dm3.chr2L 780059 68 - 23011544 AUGCAGCGCAUCCUAGGAUUUGUGGGCUCUUGUUGCUGCCGCGUCCCUUUCGAAGGACGUGCCUAUUC ..((((((((.((((.......))))....)).))))))(((((((........)))))))....... ( -21.70, z-score = -0.97, R) >droSim1.chrU 11632596 68 - 15797150 AUGCAGCGCAUCCCAGGAUUUGUGGGCUCUUGUUGCUGCCGCGUCCCUUUCGAAGGACGUGCCUAUUC ..((((((((.((((.......))))....)).))))))(((((((........)))))))....... ( -24.00, z-score = -1.63, R) >droSec1.super_14 750668 68 - 2068291 ACGCAGCGCAUCCCAGGAUUUGUGGGCUCUUGUUGCUGCCGCGUCCCUUUCGAAGGACGUGCCUAUUC ..((((((((.((((.......))))....)).))))))(((((((........)))))))....... ( -24.30, z-score = -1.60, R) >droEre2.scaffold_4929 832790 67 - 26641161 AUGCAGCGCAUCCCAGGAUUUGUGGGCUC-UGCUGCUGCCUCGUCCCCUUCGCAGGACGUGCCUAUUC ..((((((...((((.......))))...-)))))).((..(((((........))))).))...... ( -22.60, z-score = -0.85, R) >droWil1.scaffold_180772 6452536 66 + 8906247 AUCCCAUCCACUCUAUACGGUAUGAGUGUGUGUAGUAUGUCAUUGUAUUCUUUUGUACGUGCCUGU-- ....((..(((((..........)))))..))..((((((((...........)).))))))....-- ( -12.50, z-score = -0.66, R) >consensus AUGCAGCGCAUCCCAGGAUUUGUGGGCUCUUGUUGCUGCCGCGUCCCUUUCGAAGGACGUGCCUAUUC ..((((((((.((((.......))))....))).)))))(((((((........)))))))....... (-15.94 = -15.58 + -0.36)
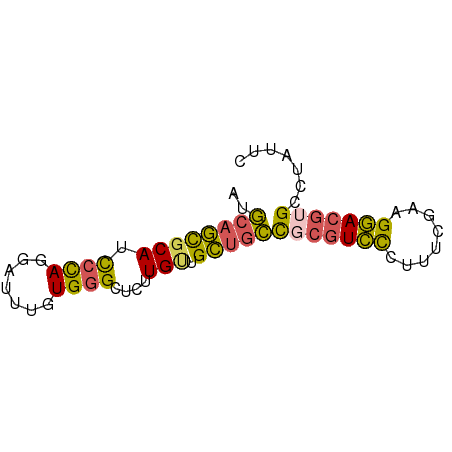
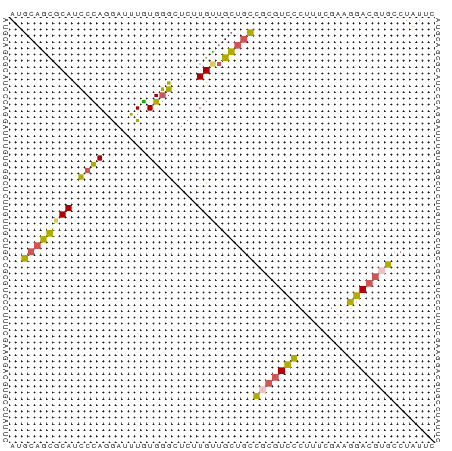
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:06:51 2011