
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,387,101 – 18,387,191 |
| Length | 90 |
| Max. P | 0.506752 |

| Location | 18,387,101 – 18,387,191 |
|---|---|
| Length | 90 |
| Sequences | 10 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 62.39 |
| Shannon entropy | 0.78487 |
| G+C content | 0.44465 |
| Mean single sequence MFE | -18.35 |
| Consensus MFE | -7.20 |
| Energy contribution | -6.93 |
| Covariance contribution | -0.27 |
| Combinations/Pair | 1.56 |
| Mean z-score | -0.66 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.506752 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
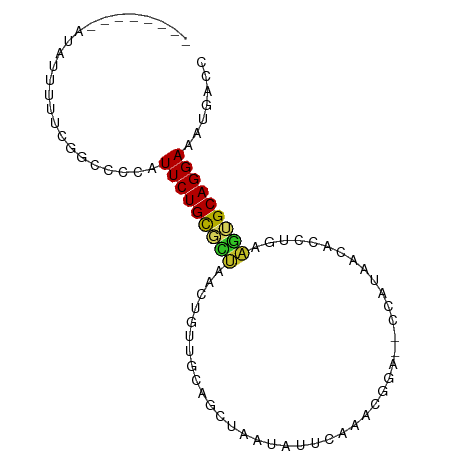
>dm3.chr3R 18387101 90 - 27905053 CCCCCUCAAGAUUUUCUGGCCCCAUUCUGCGCUAACUGUUGCAGGUAAUAUUCAAACGGAG-CCAUAACACCUGAAGUGCAGGAAAUGACC .....(((........((((.((...((((((.....).)))))((.........)))).)-))).....((((.....))))...))).. ( -18.30, z-score = 0.12, R) >droSim1.chr3R 18196786 90 - 27517382 CCCCCUCAAUAUUUUCUGGCCCCAUUCUGCGCUAACUGUUGCCGGUAAUAUUCAAACGGAG-CCAUAACACCUGAAGUGCAGGAAAUGACC .....(((........((....))(((((((((...(((((..(((....(((....))))-)).))))).....)))))))))..))).. ( -16.10, z-score = 0.49, R) >droSec1.super_0 18740240 90 - 21120651 CCCCCUCAAUAUUUUCUGGCCCCAUUCUGCGCUAACUGUUGCAGGUAAUAUUCAAACGGAG-CCAUAACACCUGAAGUGCAGGAAAUGACC .....(((........((((.((...((((((.....).)))))((.........)))).)-))).....((((.....))))...))).. ( -18.30, z-score = -0.14, R) >droYak2.chr3R 19235327 91 - 28832112 CCACCUCAAAAUUUUCUGGCCCCAUUCUGCGCUAACUGUUGCAGGUAAUAUUCAAACGGAGUUUAUAACACCUGAAGUGCAGGAAAUGACC .....(((........((....))(((((((((........(((((...((((.....)))).......))))).)))))))))..))).. ( -16.90, z-score = 0.17, R) >droEre2.scaffold_4820 780303 70 + 10470090 --------------------ACCAUUCUGCGCGAACUGUUGCAGGUAAUAUUCAAACGGAG-CCAUAACACCUGAAGUGCAGGAAAUGACC --------------------....(((((((((((.(((((....))))))))....((.(-......).))....))))))))....... ( -15.10, z-score = -0.60, R) >droAna3.scaffold_13340 21802316 86 - 23697760 ---AUAGCCAUUUUCUGAGCCACAUUCUGUUUCAACUGUUGCAGCUAAUAUUCGAAAUAG--AAACCAAAGUGUUGUAGCAGGAAAUGACU ---.....(((((((((.((.((((((((((((...(((((....)))))...)))))))--).......)))).))..)))))))))... ( -19.81, z-score = -1.59, R) >dp4.chr2 27503038 70 + 30794189 --------CCAUUUUUCAGCCCCAUUCUGCCUUGAUUGCUGCAGCUAAUAUUCA--------UGAGAGC-----AAGUGCAGGAAAUGGCU --------.........((((...((((((((((...((....)).....(((.--------...))))-----))).))))))...)))) ( -17.50, z-score = -0.08, R) >droPer1.super_7 1050489 70 - 4445127 --------CCAUUUUUCAGCCCCAUUCUGCCUUGAUUGCUGCAGCUAAUAUUCA--------UGAGAGC-----AAGUGCAGGAAAUGGCU --------.........((((...((((((((((...((....)).....(((.--------...))))-----))).))))))...)))) ( -17.50, z-score = -0.08, R) >droVir3.scaffold_13047 10113077 74 - 19223366 ---------UCAUUUUCUGGUUCGUUCUGCGUUUAUUGUUGCAGCAACAAAAAU--------CAAACAAGCGGCCAGUGCAGGAAAUGAGC ---------((((((((((....((((((((........))))).)))......--------.......((.....)).)))))))))).. ( -19.40, z-score = -1.24, R) >droGri2.scaffold_14906 6823137 72 + 14172833 ---------UCAUUUUCUGCUUUGUUCUGCGUUUAUUGUUGCAGCAAAUAGCAA--------AUAGCAA--AAACAAUGCAGGAAAUGAGC ---------(((((((((((.((((((((((........)))))......((..--------...))..--.))))).))))))))))).. ( -24.60, z-score = -3.64, R) >consensus ________AUAUUUUUCGGCCCCAUUCUGCGCUAACUGUUGCAGCUAAUAUUCAAACGGA__CCAUAACACCUGAAGUGCAGGAAAUGACC ........................(((((((((..........................................)))))))))....... ( -7.20 = -6.93 + -0.27)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:35:34 2011