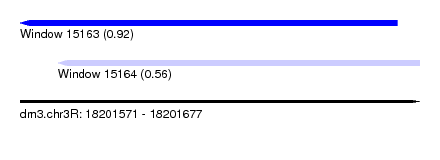

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,201,571 – 18,201,677 |
| Length | 106 |
| Max. P | 0.924825 |

| Location | 18,201,571 – 18,201,671 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 88.00 |
| Shannon entropy | 0.19537 |
| G+C content | 0.33513 |
| Mean single sequence MFE | -24.45 |
| Consensus MFE | -16.70 |
| Energy contribution | -17.70 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.924825 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
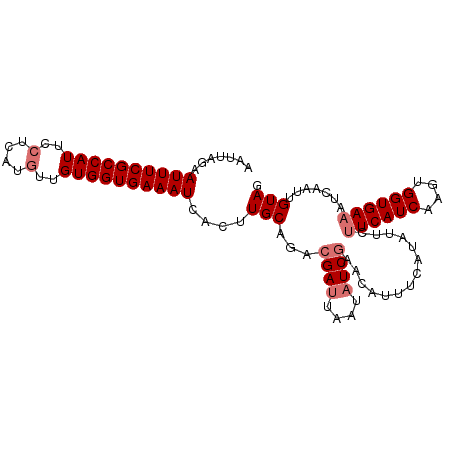
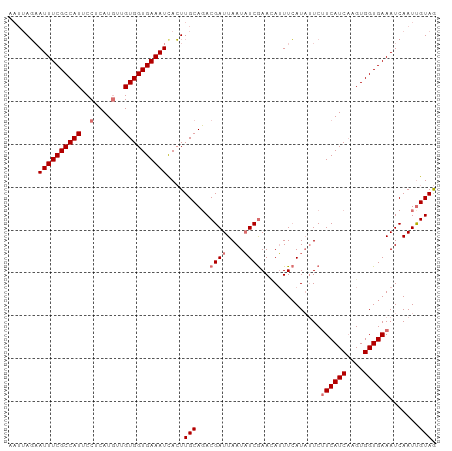
>dm3.chr3R 18201571 100 - 27905053 AAUUAGAAUUUCGCCAUUCCUCAUCAUGUGGUGAAAUCACUUGCAUACGAUUAAUAUCGAAUAUUUCAUUAUCUUCAUCAUGUGGUGAAAUCAAUUGUAG .....((.(((((((((..........)))))))))))...((((..((((....))))...((((((((((.........))))))))))....)))). ( -20.40, z-score = -1.47, R) >droYak2.chr3R 19045879 97 - 28832112 ---AACUAUUUCGCCAUUCCUCAUGUUGUGGUGAAAUCAGUUGCAGACGAUUAAUUUCAAAAAGUUGAUAUUCCUCAUCUUGUGGUGAAAUCAAUCGUAA ---((((((((((((((..(....)..)))))))))).))))....((((((.((((((..(((.(((......))).)))....)))))).)))))).. ( -29.80, z-score = -4.90, R) >droSec1.super_0 18557485 100 - 21120651 AAUUAGAAUUUCGCCAUUCCUCAUGUUGUGGUGAAAUCACUUGCAGACGAUUAAUAUCGAACAUUUCAUAUUCUUCAUCAAGUGGUGAAAUCAAUUGUAG .....((.(((((((((..(....)..)))))))))))...(((((.((((....))))...(((((((...(((....)))..)))))))...))))). ( -23.30, z-score = -2.04, R) >droSim1.chr3R 18010570 100 - 27517382 AAUUAGAAUUUCGCCAUUCCUCAUGUUGUGGUGAAAUUACUUGCAGACGAUUAAUAUCGAACAUUUCAUAUUCUUCAUCAAGUGGUGAAAUCAAUUGUAG ......(((((((((((..(....)..)))))))))))...(((((.((((....))))...(((((((...(((....)))..)))))))...))))). ( -24.30, z-score = -2.54, R) >consensus AAUUAGAAUUUCGCCAUUCCUCAUGUUGUGGUGAAAUCACUUGCAGACGAUUAAUAUCGAACAUUUCAUAUUCUUCAUCAAGUGGUGAAAUCAAUUGUAG .......((((((((((..(....)..))))))))))....(((...((((....))))..............((((((....)))))).......))). (-16.70 = -17.70 + 1.00)

| Location | 18,201,581 – 18,201,677 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 70.04 |
| Shannon entropy | 0.57815 |
| G+C content | 0.34908 |
| Mean single sequence MFE | -19.97 |
| Consensus MFE | -9.69 |
| Energy contribution | -10.22 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.25 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.563773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
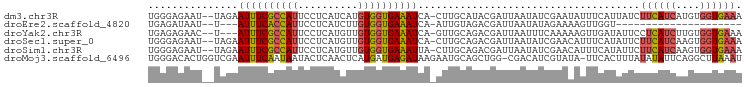
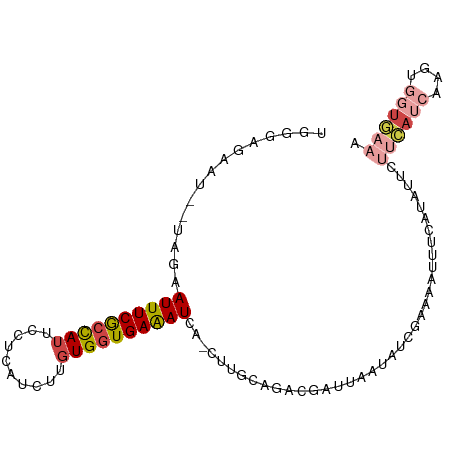
>dm3.chr3R 18201581 96 - 27905053 UGGGAGAAU--UAGAAUUUCGCCAUUCCUCAUCAUGUGGUGAAAUCA-CUUGCAUACGAUUAAUAUCGAAUAUUUCAUUAUCUUCAUCAUGUGGUGAAA (((((((..--.....)))).)))....(((((((((((((((....-........((((....)))).(((......))).))))))))))))))).. ( -23.00, z-score = -1.35, R) >droEre2.scaffold_4820 594319 72 + 10470090 UGAGAUAAU--U---AUUUCACCAUUCCUCAUCUUGUGGUGAAAUCA-AUUGUAGACGAUUAAUAUAGAAAAGUUGGU--------------------- ....(((((--(---((((((((((..........)))))))))).)-)))))...(((((..........)))))..--------------------- ( -14.60, z-score = -1.36, R) >droYak2.chr3R 19045889 93 - 28832112 UGAGAGAAC--U---AUUUCGCCAUUCCUCAUGUUGUGGUGAAAUCA-GUUGCAGACGAUUAAUUUCAAAAAGUUGAUAUUCCUCAUCUUGUGGUGAAA ......(((--(---((((((((((..(....)..)))))))))).)-)))...((..((((((((....))))))))..)).(((((....))))).. ( -22.70, z-score = -1.64, R) >droSec1.super_0 18557495 96 - 21120651 UGGGAGAAU--UAGAAUUUCGCCAUUCCUCAUGUUGUGGUGAAAUCA-CUUGCAGACGAUUAAUAUCGAACAUUUCAUAUUCUUCAUCAAGUGGUGAAA ...((((..--..((.(((((((((..(....)..))))))))))).-........((((....))))....))))......((((((....)))))). ( -22.50, z-score = -0.95, R) >droSim1.chr3R 18010580 96 - 27517382 UGGGAGAAU--UAGAAUUUCGCCAUUCCUCAUGUUGUGGUGAAAUUA-CUUGCAGACGAUUAAUAUCGAACAUUUCAUAUUCUUCAUCAAGUGGUGAAA ...((((..--...(((((((((((..(....)..))))))))))).-........((((....))))....))))......((((((....)))))). ( -23.50, z-score = -1.52, R) >droMoj3.scaffold_6496 7775451 97 + 26866924 UGGGACACUGGUCGAAUUUCAAUAAUACUCAACUCAUGAUGAGAUAAGAAUGCAGCUGG-CGACAUCGUAUA-UUCACUUUAUAUAUUCAGGCUUAAAU ((((...((((((..............((((.(....).))))........((.....)-))))...(((((-(.......)))))).))).))))... ( -13.50, z-score = 0.90, R) >consensus UGGGAGAAU__UAGAAUUUCGCCAUUCCUCAUCUUGUGGUGAAAUCA_CUUGCAGACGAUUAAUAUCGAAAAUUUCAUAUUCUUCAUCAAGUGGUGAAA ...............((((((((((..........)))))))))).....................................((((((....)))))). ( -9.69 = -10.22 + 0.53)
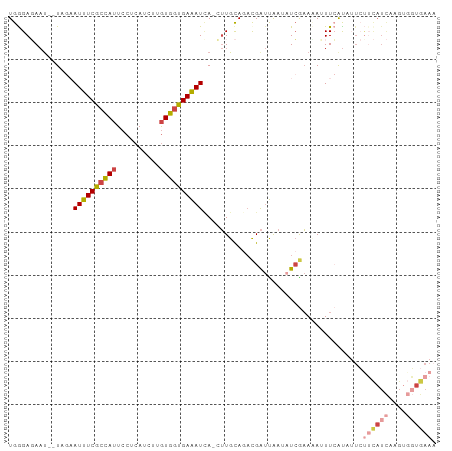
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:35:10 2011