
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,955,862 – 17,955,981 |
| Length | 119 |
| Max. P | 0.951266 |

| Location | 17,955,862 – 17,955,981 |
|---|---|
| Length | 119 |
| Sequences | 7 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 76.39 |
| Shannon entropy | 0.48806 |
| G+C content | 0.55750 |
| Mean single sequence MFE | -41.20 |
| Consensus MFE | -20.59 |
| Energy contribution | -22.67 |
| Covariance contribution | 2.08 |
| Combinations/Pair | 1.42 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.951266 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17955862 119 + 27905053 GGCAACUGCGGCAGAUGCAGCGGCAGUGCAAGGACAAGACGGACAAGUCCAACUACGAGCGAAAGCAGGCGACCGCCAAGGCGGAGGAGCUGGAACUGGAGUUGCAGUCCCGCCGUAGG .(((.((((.((.......)).)))))))..((((...........))))..((((..((....)).(((..((((....)))).((..(((.(((....))).)))..))))))))). ( -53.60, z-score = -3.67, R) >dp4.chr2 12123399 119 + 30794189 GGCAGCUGCGGCAAAUGCAGCGUCAGUGCAAGGACAAGAUGGACAAAUCCAAUUACGAACGGAAACAGGCCUCAGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAAUCGCGUCGCAAG ......((((((...((((((..((((....(..(.....)..)...((((.........(....)...((((.((....)).))))...))))))))..)))))).....)))))).. ( -41.10, z-score = -1.69, R) >droPer1.super_0 3449615 119 - 11822988 GGCAGCUGCGGCAAAUGCAGCGUCAGUGCAAGGACAAGAUGGACAAAUCCAAUUACGAACGGAAACAGGCCUCAGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAGUCCCGUCGCAAA ....((.((((....((((((..((((....(..(.....)..)...((((.........(....)...((((.((....)).))))...))))))))..))))))...)))).))... ( -42.90, z-score = -1.89, R) >droWil1.scaffold_181130 12096310 119 + 16660200 GGCAGUUAAGGCAGAUGCAGCGUCAAUGCAAAGAUAAAGUCGACAAAUGCAAUUACGAAAGGAAACAGGCUUCAGCCAAGGCUGAUGAACUUGAACAGGAGUUGCAAUCGCGUCGCAAG .((.((....)).(((((.(((....)))..................(((((((.(....)....((((..(((((....)))))....))))......)))))))...)))))))... ( -30.40, z-score = -0.66, R) >droVir3.scaffold_13047 10694126 119 - 19223366 GGCAGAUGCGGCAAAUGCAGCGUCAGUGCAAGGAUAAACUGGACAAAUCCAAUUACGAACGCAAACAGGCAACGGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAAUCACGUCGCAAA ......((((((...((((((..((((((.((......))(((....)))..........))...(((.(..((((....))))..)..)))..))))..)))))).....)))))).. ( -39.90, z-score = -2.64, R) >droGri2.scaffold_15074 1088783 119 + 7742996 GGCAGAUGCGUCAAAUGCAGCGCCAGUGCAAGGACAAACUUGACAAAUCCAAUUACGAACGCAAACAGGCAUCGGCCAAGGCUGAGGAACUGGAUCUGGAGCUGCAGUCGCGACGCAAG ......((((((...((((((.((((((((((......)))).)).(((((.........((......)).(((((....))))).....))))))))).)))))).....)))))).. ( -44.50, z-score = -3.02, R) >anoGam1.chr3L 14825420 110 + 41284009 UUACGAUCCGCAGCAUGCAGAAGACGUGCAAGGAGAAGAUGGAAACGGUUGAGAAGGAGCGGAAGCAGGC----GCUCGAGCAGGUGAGCCGG--CUGGAG---GAGUUGGCCCGCAAG ...(..(((...((((((....).)))))..)))...)..(....)(.(((((.....((....))....----.))))).)..(((.(((((--((....---.))))))).)))... ( -36.00, z-score = -1.20, R) >consensus GGCAGAUGCGGCAAAUGCAGCGUCAGUGCAAGGACAAGAUGGACAAAUCCAAUUACGAACGGAAACAGGCAUCAGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAGUCGCGUCGCAAG ......((((((...((((((.(((((....((...........................(....)......((((....)))).....))...))))).)))))).....)))))).. (-20.59 = -22.67 + 2.08)


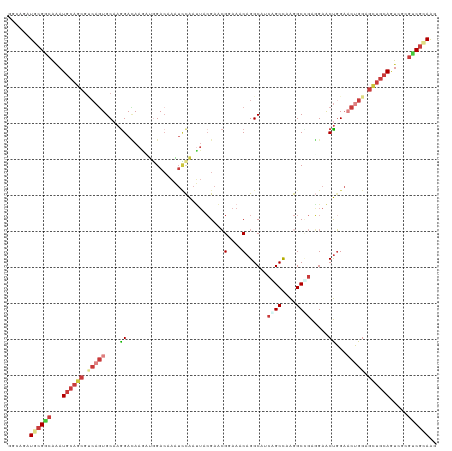
| Location | 17,955,862 – 17,955,981 |
|---|---|
| Length | 119 |
| Sequences | 7 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 76.39 |
| Shannon entropy | 0.48806 |
| G+C content | 0.55750 |
| Mean single sequence MFE | -38.31 |
| Consensus MFE | -13.68 |
| Energy contribution | -14.29 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.52 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3R 17955862 119 - 27905053 CCUACGGCGGGACUGCAACUCCAGUUCCAGCUCCUCCGCCUUGGCGGUCGCCUGCUUUCGCUCGUAGUUGGACUUGUCCGUCUUGUCCUUGCACUGCCGCUGCAUCUGCCGCAGUUGCC .((((((((((((((......)))))))(((.(..((((....))))..)...)))...)).)))))..((((..(.....)..))))..(((((((.((.......)).)))).))). ( -45.70, z-score = -2.40, R) >dp4.chr2 12123399 119 - 30794189 CUUGCGACGCGAUUGCAGCUCCAGUUCCAGUUCCUCGGCCUUGGCUGAGGCCUGUUUCCGUUCGUAAUUGGAUUUGUCCAUCUUGUCCUUGCACUGACGCUGCAUUUGCCGCAGCUGCC .(((((..((((.((((((..((((..(((..(((((((....))))))).))).........((((..((((..(.....)..))))))))))))..)))))).)))))))))..... ( -45.10, z-score = -3.26, R) >droPer1.super_0 3449615 119 + 11822988 UUUGCGACGGGACUGCAGCUCCAGUUCCAGUUCCUCGGCCUUGGCUGAGGCCUGUUUCCGUUCGUAAUUGGAUUUGUCCAUCUUGUCCUUGCACUGACGCUGCAUUUGCCGCAGCUGCC ...((..(((((.((((((..((((..(((..(((((((....))))))).))).........((((..((((..(.....)..))))))))))))..)))))).)).)))..)).... ( -42.40, z-score = -1.89, R) >droWil1.scaffold_181130 12096310 119 - 16660200 CUUGCGACGCGAUUGCAACUCCUGUUCAAGUUCAUCAGCCUUGGCUGAAGCCUGUUUCCUUUCGUAAUUGCAUUUGUCGACUUUAUCUUUGCAUUGACGCUGCAUCUGCCUUAACUGCC ....(((((((((((((((....)))((.(((..(((((....)))))))).)).........))))))))....))))...........(((((((.((.......)).)))).))). ( -26.60, z-score = -0.78, R) >droVir3.scaffold_13047 10694126 119 + 19223366 UUUGCGACGUGAUUGCAGCUCCAGUUCCAGUUCCUCGGCCUUGGCCGUUGCCUGUUUGCGUUCGUAAUUGGAUUUGUCCAGUUUAUCCUUGCACUGACGCUGCAUUUGCCGCAUCUGCC ..((((..(..(.((((((..((((..(((.....((((....))))....)))...(((.....(((((((....)))))))......)))))))..)))))).)..)))))...... ( -40.20, z-score = -3.01, R) >droGri2.scaffold_15074 1088783 119 - 7742996 CUUGCGUCGCGACUGCAGCUCCAGAUCCAGUUCCUCAGCCUUGGCCGAUGCCUGUUUGCGUUCGUAAUUGGAUUUGUCAAGUUUGUCCUUGCACUGGCGCUGCAUUUGACGCAUCUGCC ..(((((((....((((((.((((((((((((.....((....))((((((......))).))).)))))))).((.((((......)))))))))).))))))..)))))))...... ( -42.20, z-score = -2.33, R) >anoGam1.chr3L 14825420 110 - 41284009 CUUGCGGGCCAACUC---CUCCAG--CCGGCUCACCUGCUCGAGC----GCCUGCUUCCGCUCCUUCUCAACCGUUUCCAUCUUCUCCUUGCACGUCUUCUGCAUGCUGCGGAUCGUAA ...((((((......---(((.((--(.((....)).))).))).----)))))).....................................(((...(((((.....))))).))).. ( -26.00, z-score = -0.38, R) >consensus CUUGCGACGCGACUGCAGCUCCAGUUCCAGUUCCUCGGCCUUGGCUGAGGCCUGUUUCCGUUCGUAAUUGGAUUUGUCCAUCUUGUCCUUGCACUGACGCUGCAUUUGCCGCAGCUGCC .((((((((((((((.(((....))).))).....((((....))))........)))))).)))))......................((((.......))))............... (-13.68 = -14.29 + 0.61)

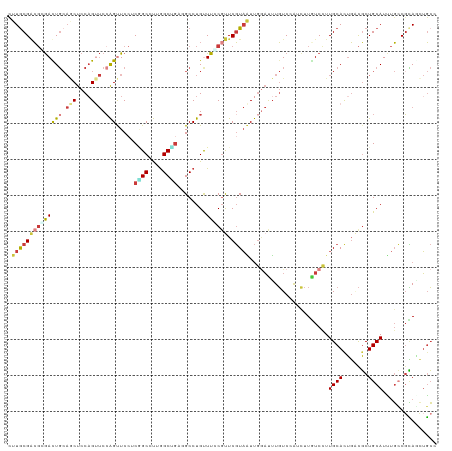
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:34:16 2011