
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,922,246 – 17,922,344 |
| Length | 98 |
| Max. P | 0.557995 |

| Location | 17,922,246 – 17,922,344 |
|---|---|
| Length | 98 |
| Sequences | 8 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 89.31 |
| Shannon entropy | 0.21262 |
| G+C content | 0.37275 |
| Mean single sequence MFE | -23.26 |
| Consensus MFE | -14.86 |
| Energy contribution | -15.41 |
| Covariance contribution | 0.55 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.557995 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17922246 98 - 27905053 CAUAAUCACAAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUUAGCUGCACAAAGUCAAAGUCGUUGACGUCGUCAU----UCUCGAUGACUCAAA ..............(((((((.((((((((-.((....)).))))))))....)).)))))(((((.....)))))((((((..----....))))))..... ( -22.00, z-score = -2.08, R) >droSim1.chrU 7294763 95 + 15797150 CGUGA--GCGAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUU-GCUGCACAAAGUCAAAGUCGUUGACGUCGUCAU----UCUCGAUGACUCAAA .((.(--((((...((((.......(((((-((((...(((((....)))-))))).)))))))))).))))).))((((((..----....))))))..... ( -28.31, z-score = -2.25, R) >droSec1.super_0 18290961 97 - 21120651 CAUAAUCACAAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUU-GCUGCACAAAGUCAAAGUCGUUGACGUCGUCAU----UCUCGAUGACUCAAA ..............(((((((.((((((((-.((....)).)))))))).-..)).)))))(((((.....)))))((((((..----....))))))..... ( -22.60, z-score = -2.01, R) >droYak2.chr3R 18762999 97 - 28832112 CAUAAUCACAAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUU-GCUGCACAAAGUCAAAGUCGUUGACGUCGUCAU----UCUCGAUGACUCAAA ..............(((((((.((((((((-.((....)).)))))))).-..)).)))))(((((.....)))))((((((..----....))))))..... ( -22.60, z-score = -2.01, R) >droEre2.scaffold_4820 315110 97 + 10470090 CAUAAUCACAAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUU-GCUGCACAAAGUCAAAGUCGUUGACGUCGUCAU----UCUCGAUGACUCAAA ..............(((((((.((((((((-.((....)).)))))))).-..)).)))))(((((.....)))))((((((..----....))))))..... ( -22.60, z-score = -2.01, R) >droAna3.scaffold_13340 8928556 96 - 23697760 CAUAAUCACAAAAAUUUGUCAUAAAACUUUCGACAGACGCCAAAGUUUUUGGCUGCACAAAGUCAAAGACAU---CGCCAUCUU----UCUCGAUGGCUUAAA ..............((((((...........)))))).......((((((((((......))))))))))..---.((((((..----....))))))..... ( -25.60, z-score = -4.37, R) >dp4.chr2 12086307 102 - 30794189 CAUAAUCACAAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUGCGUUGCACAAAGUCAAAGUCGUUGACGUCGUCAUCAAAUUUCAAUGCUAUAGA ...........(((((((.......(((((-(((((.(((((.....))))))))).))))))....(.((....)).).....)))))))............ ( -20.00, z-score = -1.01, R) >droPer1.super_0 3412337 102 + 11822988 CAUAAUCACAAAAAUUUGUCAUAAAACUUU-GGCAGACGCAAAAGUUUUGCGCUGCACAAAGUCAAAGUCGUUGACGUCGUCAUCAAAUUUCAAUGCUAUAGA ...........(((((((.......(((((-(((((.(((((.....))))))))).))))))....(.((....)).).....)))))))............ ( -22.40, z-score = -1.76, R) >consensus CAUAAUCACAAAAAUUUGUCAUAAAACUUU_GGCAGACGCAAAAGUUUUU_GCUGCACAAAGUCAAAGUCGUUGACGUCGUCAU____UCUCGAUGACUCAAA ..............(((((((.(((((((...((....))..)))))))....)).)))))(((((.....)))))((((((..........))))))..... (-14.86 = -15.41 + 0.55)


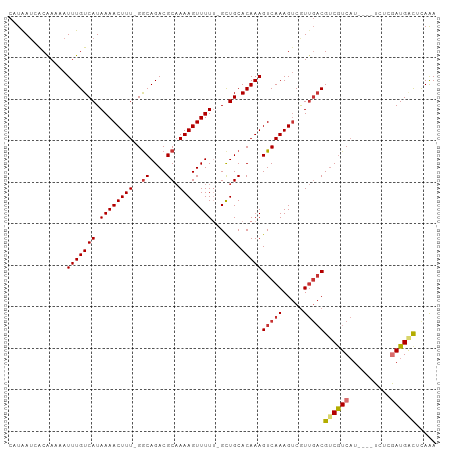
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:34:10 2011