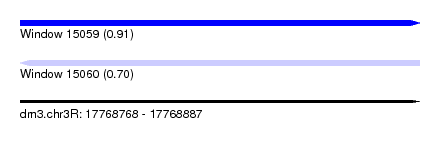

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,768,768 – 17,768,887 |
| Length | 119 |
| Max. P | 0.906716 |

| Location | 17,768,768 – 17,768,887 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 97.20 |
| Shannon entropy | 0.03858 |
| G+C content | 0.48877 |
| Mean single sequence MFE | -40.53 |
| Consensus MFE | -37.82 |
| Energy contribution | -38.60 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.02 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.93 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.906716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
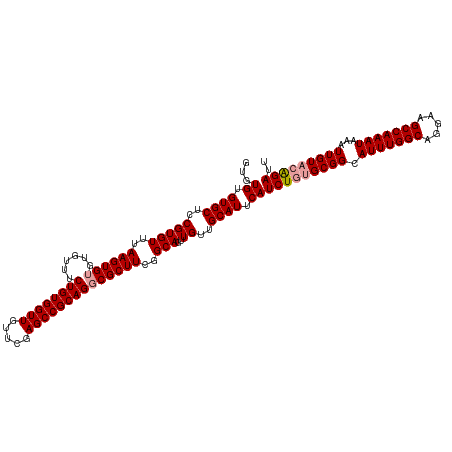
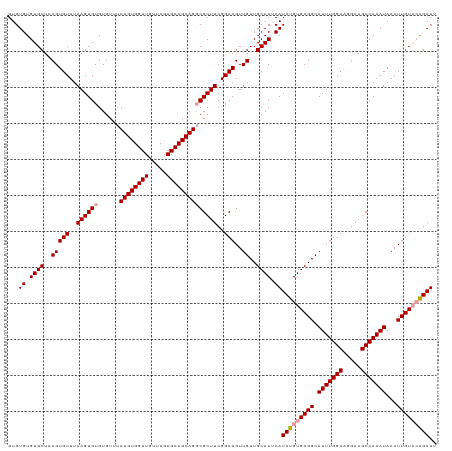
>dm3.chr3R 17768768 119 + 27905053 GUGUGUGUGCUCCGUGUUUAAGUGUGUGUUUCUGUGGUUGUUCGAGCCGCAGGCGCUUCGGCAGUUGUUGCAUUCAUCUGUGCGGCAUUUGGCAGGAAGCCAAAUAAAUUGUAGAGAUU ......((((..((..(......)..))...((((((((.....))))))))))))(((.(((((((((((((......)))))))(((((((.....))))))).)))))).)))... ( -38.10, z-score = -1.32, R) >droSim1.chr3R 17579837 119 + 27517382 GUGUGUGUGCUGCGUGUUUAAGUGUGUGUUUCUGUGGUUGUUCGAGCCGCAGGCGCUUCGGCAGUUGUUGCAUUCAUCUGUGCGGCAUUUGGCAGGAAGCCAAAUAAAUUGUACAGAUU ...((.((((.((((((..((((((......((((((((.....))))))))))))))..)))..))).)))).))(((((((((.(((((((.....)))))))...))))))))).. ( -43.30, z-score = -2.22, R) >droSec1.super_0 18139281 118 + 21120651 GUGUGUGUGCUCCGUGUUUAAGUGUGUGUUUCUGUGGUUGUUCGAGCCGCAGUCGCUUCGGCAGUUGUUGCAUUCAUCUGUGCGGCAUUUGGCAGGAAGCCAAAUAAAUUGU-CGGAUU .........................(((...((((((((.....)))))))).)))(((((((((((((((((......)))))))(((((((.....))))))).))))))-)))).. ( -40.20, z-score = -2.46, R) >consensus GUGUGUGUGCUCCGUGUUUAAGUGUGUGUUUCUGUGGUUGUUCGAGCCGCAGGCGCUUCGGCAGUUGUUGCAUUCAUCUGUGCGGCAUUUGGCAGGAAGCCAAAUAAAUUGUACAGAUU ...((.((((..(((((..((((((......((((((((.....))))))))))))))..)))..))..)))).))(((((((((.(((((((.....)))))))...))))))))).. (-37.82 = -38.60 + 0.78)

| Location | 17,768,768 – 17,768,887 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 97.20 |
| Shannon entropy | 0.03858 |
| G+C content | 0.48877 |
| Mean single sequence MFE | -25.54 |
| Consensus MFE | -21.66 |
| Energy contribution | -22.44 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.697622 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
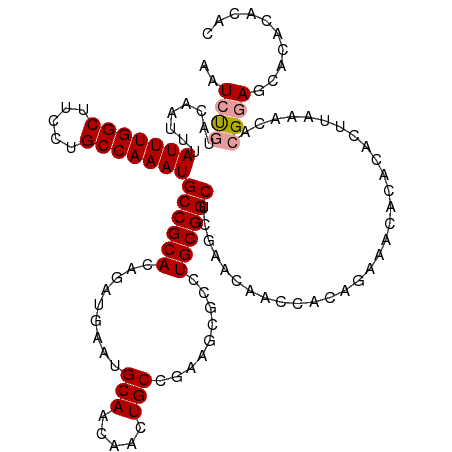
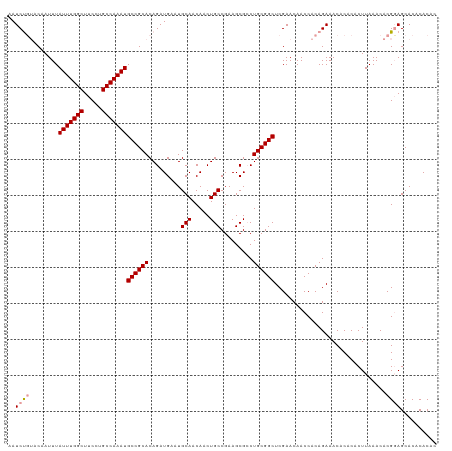
>dm3.chr3R 17768768 119 - 27905053 AAUCUCUACAAUUUAUUUGGCUUCCUGCCAAAUGCCGCACAGAUGAAUGCAACAACUGCCGAAGCGCCUGCGGCUCGAACAACCACAGAAACACACACUUAAACACGGAGCACACACAC ...((((.......(((((((.....)))))))((((((.........(((.....))).........))))))................................))))......... ( -23.87, z-score = -1.60, R) >droSim1.chr3R 17579837 119 - 27517382 AAUCUGUACAAUUUAUUUGGCUUCCUGCCAAAUGCCGCACAGAUGAAUGCAACAACUGCCGAAGCGCCUGCGGCUCGAACAACCACAGAAACACACACUUAAACACGCAGCACACACAC ..(((((.......(((((((.....)))))))((((((.........(((.....))).........))))))..........))))).............................. ( -26.17, z-score = -1.75, R) >droSec1.super_0 18139281 118 - 21120651 AAUCCG-ACAAUUUAUUUGGCUUCCUGCCAAAUGCCGCACAGAUGAAUGCAACAACUGCCGAAGCGACUGCGGCUCGAACAACCACAGAAACACACACUUAAACACGGAGCACACACAC ..((((-.......(((((((.....)))))))((((((.........(((.....))).........))))))...............................)))).......... ( -26.57, z-score = -2.83, R) >consensus AAUCUGUACAAUUUAUUUGGCUUCCUGCCAAAUGCCGCACAGAUGAAUGCAACAACUGCCGAAGCGCCUGCGGCUCGAACAACCACAGAAACACACACUUAAACACGGAGCACACACAC ..(((((.......(((((((.....)))))))((((((.........(((.....))).........))))))..............................))))).......... (-21.66 = -22.44 + 0.78)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:33:44 2011