| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,631,641 – 17,631,742 |
| Length | 101 |
| Max. P | 0.996216 |

| Location | 17,631,641 – 17,631,742 |
|---|---|
| Length | 101 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 71.61 |
| Shannon entropy | 0.55836 |
| G+C content | 0.42967 |
| Mean single sequence MFE | -25.81 |
| Consensus MFE | -10.69 |
| Energy contribution | -12.21 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.90 |
| SVM RNA-class probability | 0.996216 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17631641 101 + 27905053 ------GCAGGCGAAGGCGGAACGGGGCGUUUUAUCUAAACGGUU--CCAAAAGGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUAAAAAUAAAAACGGCGGCAGGGGAG-- ------((..((....)).....(((((.((((..((((((((((--((....)).))))))))))..)))).)))))...................))..........-- ( -28.20, z-score = -2.64, R) >droEre2.scaffold_4820 16475 101 - 10470090 ------GCAGGCGGAGGCGGAACAGGGCUUUUUAUCUAAACGGUU--CCAAAAGGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUAAAAAUAAAAACGGCGG-CAGGGGAA- ------((.(.((..(......)((((((((((..((((((((((--((....)).))))))))))..)))))))))).................)).).)-).......- ( -31.90, z-score = -3.92, R) >droYak2.chr3R 18452375 101 + 28832112 ------ACAGGCGAAGGCGGAACGGGGCUGUUUAUCUAAACGGUU--CCAAAAGGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUAAAAAUAAAAACGGCGG-CAGGGGGA- ------....((....)).....((((((.(((..((((((((((--((....)).))))))))))..))).)))))).......................-........- ( -26.60, z-score = -1.98, R) >droSec1.super_0 18004024 103 + 21120651 ------GCAGGCGAAGGCGGAGCAGGGCUUUUUAUCUAAACGGUU--CCAAAAGGCAAUCGUUUGGGGAAAAAGUCUUAAAAAUAAAAAUAAAAACGGCGGGCAGGGGGAG ------((..((....)).....((((((((((..((((((((((--((....)).))))))))))..))))))))))...................))............ ( -29.70, z-score = -3.79, R) >droSim1.chr3R 17437316 101 + 27517382 ------GCAGGCGAAGGCGGAGUAGGGCUUUUUAUCUAAACGGUU--CCAAAAGGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUAAAAAUAAAAACGGCGGCAGGGGAG-- ------((..((....))....(((((((((((..((((((((((--((....)).))))))))))..)))))))))))..................))..........-- ( -33.70, z-score = -4.62, R) >droPer1.super_0 313642 90 + 11822988 -------CAGAGAUACGGAG--GAGGGCU-UUUAUCUAAACGGCU--CCAAACAGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUGUAAAUAAAAAGAACAAC--------- -------......((((...--.((((((-(((.(((((((((((--......)))...))))))))..))))))))).....))))...............--------- ( -23.10, z-score = -3.65, R) >dp4.chr2 6488525 92 + 30794189 -------CAGAGAUACGGAGAGGAGGGCU-UUUAUCUAAACGGCU--CCAAACAGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUGUAAAUAAAAAGAACAAC--------- -------......((((......((((((-(((.(((((((((((--......)))...))))))))..))))))))).....))))...............--------- ( -22.90, z-score = -3.37, R) >droMoj3.scaffold_6540 17216257 106 + 34148556 AUAUUUUCAUCCACAGGCCAAUUGAGACUUUUAAUUAAACCAAACGGCCAAAUUACAGAAGACUGCCUAGCCAAC---AAAAAUAUAAUUUUAUGCGCCUAACAGAUAA-- ........(((...((((((((((((...))))))).........(((.......(((....)))....)))...---................).))))....)))..-- ( -10.40, z-score = 0.27, R) >consensus ______GCAGGCGAAGGCGGAACAGGGCUUUUUAUCUAAACGGUU__CCAAAAGGCAAUCGUUUGGGGAAAAAGCCUUAAAAAUAAAAAUAAAAACGGCGGCCAGGGAG__ .......................((((((.(((.(((((((((...............))))))))).))).))))))................................. (-10.69 = -12.21 + 1.52)
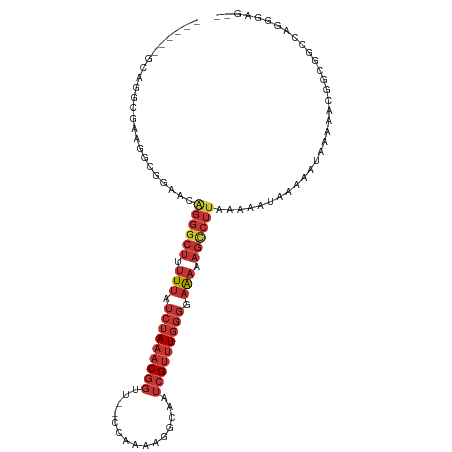


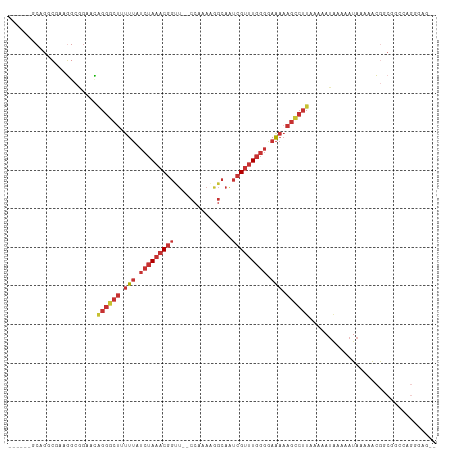
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:33:20 2011