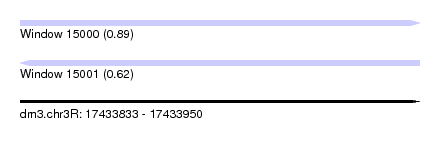

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,433,833 – 17,433,950 |
| Length | 117 |
| Max. P | 0.887957 |

| Location | 17,433,833 – 17,433,950 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 90.37 |
| Shannon entropy | 0.17433 |
| G+C content | 0.37371 |
| Mean single sequence MFE | -30.62 |
| Consensus MFE | -23.60 |
| Energy contribution | -23.64 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.887957 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
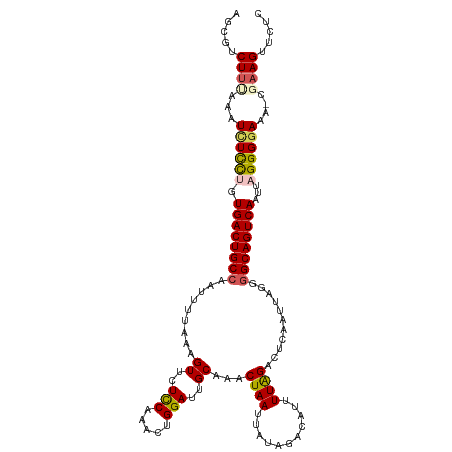
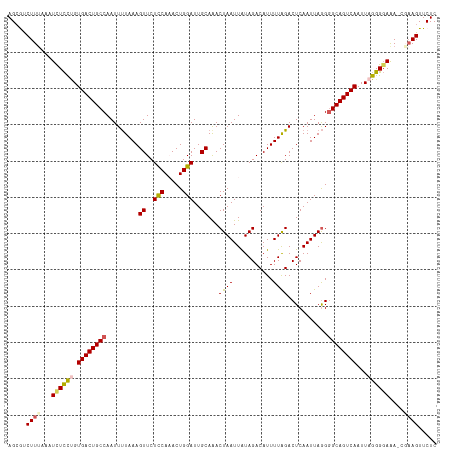
>dm3.chr3R 17433833 117 + 27905053 AGCGUCUUCGAAUUUCCUGUGACUGCCAAUUUUAAAGUUCUCCAAACUGGAUUGCAAACUAAUUAUAGACAUUUUGGACUCAAUUAGGGGCAGUCAAUUAGGGGAAA-CGAAGUUCUC .....(((((..(..((((((((((((.........((..(((.....)))..))...((((((.....(.....).....)))))).))))))))..))))..)..-)))))..... ( -34.10, z-score = -2.97, R) >droSim1.chr3R 23482843 118 - 27517382 AGCGUCUUUAAAUCUCCUGUGACUGCCAAUUUUAAAGUUCUCCACACUGGAUUGCAAACUAAUUAUAGACAUUUUAGACUCAAUUAGGGGCAGUCAAUUAGGGGAAAACGAAGUUCUC .....((((...(((((((((((((((.........((..(((.....)))..))...((((((..((.(......).)).)))))).))))))))..)))))))....))))..... ( -30.70, z-score = -2.39, R) >droSec1.super_6 302994 117 - 4358794 AGCGUCUUUAAAUCUCCCGUGACUGCCAAUUUUAAAGUUCUCCACACUGGAUUGCAAACUAAUUAUAGACAUUUUAGACUCAAUUAGGGGCAGUCAAUUAGGGGAAA-CUAAGUUCUC .....(((....(((((..((((((((.........((..(((.....)))..))...((((((..((.(......).)).)))))).))))))))....)))))..-..)))..... ( -28.60, z-score = -2.01, R) >droYak2.chr3R 6274280 114 + 28832112 AGCAUCUUUUUAUCUUUUGUGACUGCCAAUUCUAAAGUUCUUCAAACUGGAUUGCAAACUAAUUGUAGACAUUUUAGACUCAAUUAAGUGCAGUCAAUUAGGGUAGA----AGUUCUC ......((((((((((...(((((((((((((...((((.....))))))))))...((((((((.((.(......).))))))).))))))))))...))))))))----))..... ( -26.40, z-score = -1.71, R) >droEre2.scaffold_4770 13517885 117 - 17746568 AGGACCUUUUAAUCUCCAGUGACUGCCAAUUUUAAAGUUCUCCAAACUGGAUUGCAGACUAAUUAUAGACAUUUUAGACUCAAUUAGGGGCAGUCAAUUAGGGGAAA-AGAAGUUCUC .(((.(((((..(((((..((((((((.........((..(((.....)))..))...((((((..((.(......).)).)))))).))))))))....)))))..-))))).))). ( -33.30, z-score = -2.57, R) >consensus AGCGUCUUUAAAUCUCCUGUGACUGCCAAUUUUAAAGUUCUCCAAACUGGAUUGCAAACUAAUUAUAGACAUUUUAGACUCAAUUAGGGGCAGUCAAUUAGGGGAAA_CGAAGUUCUC ............((((((.((((((((.........((..(((.....)))..))...((((...........))))...........))))))))...))))))............. (-23.60 = -23.64 + 0.04)

| Location | 17,433,833 – 17,433,950 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 90.37 |
| Shannon entropy | 0.17433 |
| G+C content | 0.37371 |
| Mean single sequence MFE | -27.04 |
| Consensus MFE | -21.54 |
| Energy contribution | -21.90 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.622919 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
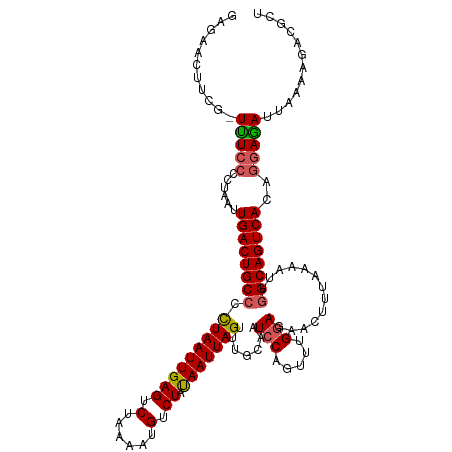
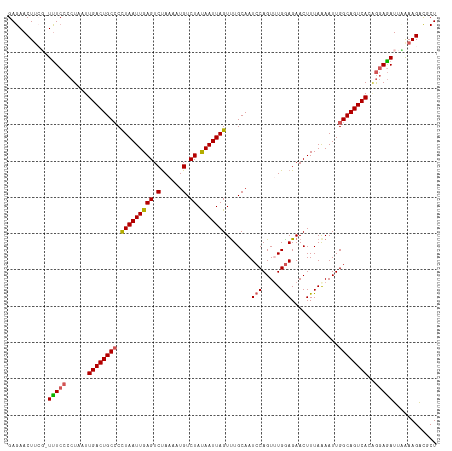
>dm3.chr3R 17433833 117 - 27905053 GAGAACUUCG-UUUCCCCUAAUUGACUGCCCCUAAUUGAGUCCAAAAUGUCUAUAAUUAGUUUGCAAUCCAGUUUGGAGAACUUUAAAAUUGGCAGUCACAGGAAAUUCGAAGACGCU .....(((((-..(..(((...((((((((.(((((((((.(......).)).))))))).......(((.....))).............)))))))).)))..)..)))))..... ( -29.50, z-score = -2.53, R) >droSim1.chr3R 23482843 118 + 27517382 GAGAACUUCGUUUUCCCCUAAUUGACUGCCCCUAAUUGAGUCUAAAAUGUCUAUAAUUAGUUUGCAAUCCAGUGUGGAGAACUUUAAAAUUGGCAGUCACAGGAGAUUUAAAGACGCU ........(((((((.(((...((((((((.(((((((((.(......).)).)))))))((..((......))..)).............)))))))).))).)).....))))).. ( -27.00, z-score = -1.53, R) >droSec1.super_6 302994 117 + 4358794 GAGAACUUAG-UUUCCCCUAAUUGACUGCCCCUAAUUGAGUCUAAAAUGUCUAUAAUUAGUUUGCAAUCCAGUGUGGAGAACUUUAAAAUUGGCAGUCACGGGAGAUUUAAAGACGCU .....(((((-((((((.....((((((((.(((((((((.(......).)).)))))))((..((......))..)).............)))))))).))))))))..)))..... ( -30.60, z-score = -2.49, R) >droYak2.chr3R 6274280 114 - 28832112 GAGAACU----UCUACCCUAAUUGACUGCACUUAAUUGAGUCUAAAAUGUCUACAAUUAGUUUGCAAUCCAGUUUGAAGAACUUUAGAAUUGGCAGUCACAAAAGAUAAAAAGAUGCU .......----...........(((((((..(((((((((.(......).)).))))))).........(((((((((....))))).)))))))))))................... ( -19.30, z-score = -0.11, R) >droEre2.scaffold_4770 13517885 117 + 17746568 GAGAACUUCU-UUUCCCCUAAUUGACUGCCCCUAAUUGAGUCUAAAAUGUCUAUAAUUAGUCUGCAAUCCAGUUUGGAGAACUUUAAAAUUGGCAGUCACUGGAGAUUAAAAGGUCCU ..((.(((.(-..((.((....((((((((.(((((((((.(......).)).)))))))(((.(((......))).)))...........))))))))..)).))..).))).)).. ( -28.80, z-score = -2.04, R) >consensus GAGAACUUCG_UUUCCCCUAAUUGACUGCCCCUAAUUGAGUCUAAAAUGUCUAUAAUUAGUUUGCAAUCCAGUUUGGAGAACUUUAAAAUUGGCAGUCACAGGAGAUUAAAAGACGCU .....(((...(((((......((((((((.(((((((((.(......).)).))))))).......(((.....))).............))))))))..)))))....)))..... (-21.54 = -21.90 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:55 2011