
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,369,838 – 17,369,893 |
| Length | 55 |
| Max. P | 0.898419 |

| Location | 17,369,838 – 17,369,893 |
|---|---|
| Length | 55 |
| Sequences | 6 |
| Columns | 55 |
| Reading direction | forward |
| Mean pairwise identity | 66.99 |
| Shannon entropy | 0.63901 |
| G+C content | 0.39853 |
| Mean single sequence MFE | -10.75 |
| Consensus MFE | -5.49 |
| Energy contribution | -5.77 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.33 |
| Mean z-score | -0.91 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.801457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
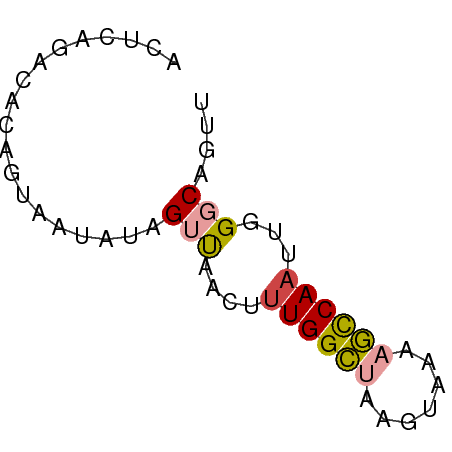
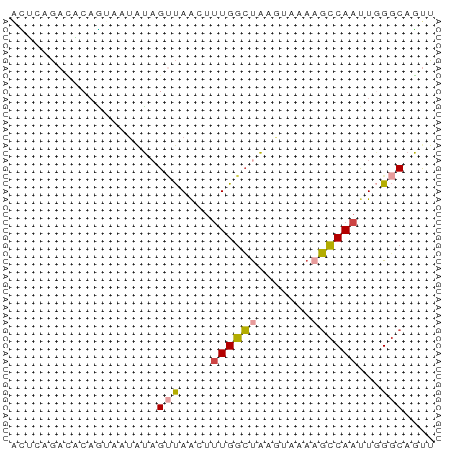
>dm3.chr3R 17369838 55 + 27905053 ACUCAGACACAGUCAUAUAGGUAACUUUGGCUAAGUAAAAGCCAAUUCGGCAGUU .....(((..(((((...((....)).)))))........(((.....))).))) ( -10.00, z-score = -1.19, R) >droSim1.chr3R 23417208 55 - 27517382 ACUCAGACACAGUCAUAUAGUUAACUUUGGCUAAGUAAAAGCCAAUUGGGCAGCU .....(((...)))....((((..((((((((.......))))))..))..)))) ( -10.30, z-score = -0.48, R) >droSec1.super_6 238717 54 - 4358794 ACUCAGACACAGUC-UAUAGUUAACUUUGGCUAAGUAAAAGCCAAUUGGGCAGUU ....((((...)))-)...(((....((((((.......))))))...))).... ( -10.50, z-score = -0.63, R) >droYak2.chr3R 6204642 54 + 28832112 AUACUGAUUCAAGAUUGUUGUUAGGUCUGGCUAAA-AAAAGCCAAUUGGGCAGUU ..((((.(((((((((.......))))(((((...-...))))).))))))))). ( -12.90, z-score = -1.53, R) >droEre2.scaffold_4770 13450038 53 - 17746568 AUACUGAUUCAGGAUUGUAGUUAAGUUUGGCUAGA-AAA-GCCAAUUGGGCAGUU ..((((.((((((((....)))....((((((...-..)-)))))))))))))). ( -10.90, z-score = -0.86, R) >droAna3.scaffold_13340 17707958 55 + 23697760 GACCUACGGCCUUAAAAAAGCCAAACUUGGGCUUGAGAACUUCAACUUGCCAUCU .......(((.......(((((.......)))))(((........)))))).... ( -9.90, z-score = -0.79, R) >consensus ACUCAGACACAGUAAUAUAGUUAACUUUGGCUAAGUAAAAGCCAAUUGGGCAGUU ...................(((....((((((.......))))))...))).... ( -5.49 = -5.77 + 0.28)

| Location | 17,369,838 – 17,369,893 |
|---|---|
| Length | 55 |
| Sequences | 6 |
| Columns | 55 |
| Reading direction | reverse |
| Mean pairwise identity | 66.99 |
| Shannon entropy | 0.63901 |
| G+C content | 0.39853 |
| Mean single sequence MFE | -11.52 |
| Consensus MFE | -4.62 |
| Energy contribution | -5.15 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.898419 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
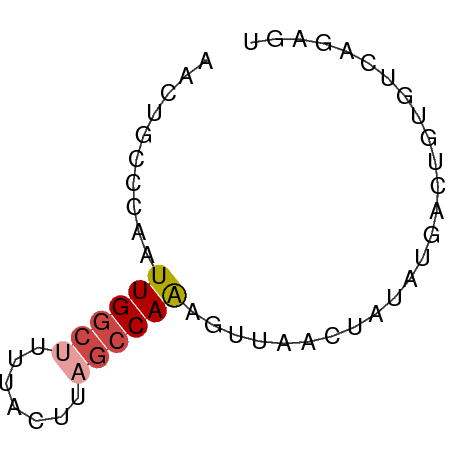
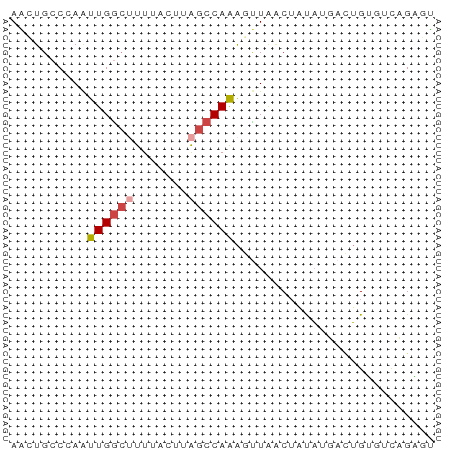
>dm3.chr3R 17369838 55 - 27905053 AACUGCCGAAUUGGCUUUUACUUAGCCAAAGUUACCUAUAUGACUGUGUCUGAGU ....(((.....)))....((((((.((.(((((......))))).)).)))))) ( -13.10, z-score = -2.37, R) >droSim1.chr3R 23417208 55 + 27517382 AGCUGCCCAAUUGGCUUUUACUUAGCCAAAGUUAACUAUAUGACUGUGUCUGAGU ....(((.....)))....((((((.((.(((((......))))).)).)))))) ( -13.00, z-score = -1.52, R) >droSec1.super_6 238717 54 + 4358794 AACUGCCCAAUUGGCUUUUACUUAGCCAAAGUUAACUAUA-GACUGUGUCUGAGU ....(((.....)))....((((((.((.((((.......-)))).)).)))))) ( -11.70, z-score = -1.36, R) >droYak2.chr3R 6204642 54 - 28832112 AACUGCCCAAUUGGCUUUU-UUUAGCCAGACCUAACAACAAUCUUGAAUCAGUAU .((((.....((((((...-...))))))......(((.....)))...)))).. ( -9.50, z-score = -1.82, R) >droEre2.scaffold_4770 13450038 53 + 17746568 AACUGCCCAAUUGGC-UUU-UCUAGCCAAACUUAACUACAAUCCUGAAUCAGUAU .((((.....(((((-(..-...)))))).........((....))...)))).. ( -9.00, z-score = -2.28, R) >droAna3.scaffold_13340 17707958 55 - 23697760 AGAUGGCAAGUUGAAGUUCUCAAGCCCAAGUUUGGCUUUUUUAAGGCCGUAGGUC ..(((((...((((((...((((((....))))))...)))))).)))))..... ( -12.80, z-score = -0.62, R) >consensus AACUGCCCAAUUGGCUUUUACUUAGCCAAAGUUAACUAUAUGACUGUGUCAGAGU ..........((((((.......)))))).......................... ( -4.62 = -5.15 + 0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:36 2011