
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,365,364 – 17,365,454 |
| Length | 90 |
| Max. P | 0.558930 |

| Location | 17,365,364 – 17,365,454 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 65.07 |
| Shannon entropy | 0.60841 |
| G+C content | 0.42308 |
| Mean single sequence MFE | -16.70 |
| Consensus MFE | -6.65 |
| Energy contribution | -6.34 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.558930 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17365364 90 - 27905053 GACAGUUUUUGGCAGAAGACAACAAACGGAGGGCCC-GACAAACAUCGCAGG-------------GAUAUAAAUGUCAGCGAGUCAAUGCUGAACAAACAAAAA ....(((((..(((...(((......(((.....))-).......((((...-------------((((....)))).)))))))..)))..)..))))..... ( -21.00, z-score = -2.13, R) >droSim1.chrU 7261376 78 - 15797150 -----------GAGAUAGAUGAGGACACGAGACCC--GACAAACAUCGCAGG-------------CAUAUAAAUGUCAGCGAGUCAAUGCUGAACAAACAAAUA -----------....(((.....(((.((.....)--).......((((.((-------------(((....))))).)))))))....)))............ ( -14.60, z-score = -1.62, R) >droSec1.super_6 234182 91 + 4358794 GACAGAUUUUGGCAGGAGACAACAAACGGAGGGCCCGGACAAACAUCGCAGG-------------CAUAUAAAUGUCAGCGAGUCAAUGCUGAACAAACAAAUA ..(((...(((((.(....)......(((.....)))........((((.((-------------(((....))))).)))))))))..)))............ ( -23.40, z-score = -2.97, R) >droYak2.chr3R 6194791 103 - 28832112 GACAGAUUUUGGCAGGAGACAACAAAAGGAGAGGCA-AACAAACAUCGCAGGGAUAUAAAUGUCAGAUAUAAAUGUCAGCGAGUCAAUGCUGAACAAACAAAUA ..(((...(((((.(....)................-........((((...((((....)))).((((....)))).)))))))))..)))............ ( -19.30, z-score = -2.29, R) >droEre2.scaffold_4770 13445300 90 + 17746568 GACAGAUUUUGGCAGGGGACAACAAACGGCGAGCCA-GACAAACAUCGAAGG-------------GAUAUAAAUGUCAGCGAGUCAAUGCUGAACAAACAAAUA .......((..(((...(((.......((....)).-........(((....-------------((((....))))..))))))..)))..)).......... ( -17.40, z-score = -1.56, R) >droAna3.scaffold_13340 17703573 71 - 23697760 -------------GACAGAUUUGGACAGGAGUC----AGUAAACA---GAGG-------------GGCAUAAAUGUCAGCAAGUCAAUGCUGAACAAACAAAUG -------------.....((((((((....)))----........---....-------------........((((((((......))))).)))..))))). ( -13.10, z-score = -1.15, R) >droVir3.scaffold_13047 4670312 76 + 19223366 -------------GACAGAUUUUAGUAUUGAGCA---AACAGACUUCAAAGGG------------UGCAAAAAUGUCAGCGAGUCAAUGCUGAGCAAACAAAUA -------------.......((((((((((((((---..(..........)..------------))).....((....))..))))))))))).......... ( -13.20, z-score = -0.05, R) >droMoj3.scaffold_6540 16151708 76 + 34148556 -------------GACAGAUUUGCGCAUUGUACA---AACACACUUCAAAGGG------------CACAAAAAUGUGAGUGGCUCAAUGCUGAGCAAACAAAUA -------------......(((((((((((....---..(((.((.....)).------------((((....)))).)))...))))))...)))))...... ( -14.80, z-score = -0.09, R) >droGri2.scaffold_14830 5416018 76 - 6267026 -------------GACAGAGUUUGGCAUUGGGCA---AACAAACUUCAAAGGG------------UACAAAAAUGUCAGCGAGUCACUGCUGAGCAAACAAAUA -------------......(((((.(....).))---))).............------------........((((((((......)))))).))........ ( -13.50, z-score = 0.16, R) >consensus ___________G_GACAGACAACAAAAGGAGACCC__AACAAACAUCGAAGG_____________GAUAUAAAUGUCAGCGAGUCAAUGCUGAACAAACAAAUA .........................................................................((((((((......))))).)))........ ( -6.65 = -6.34 + -0.31)

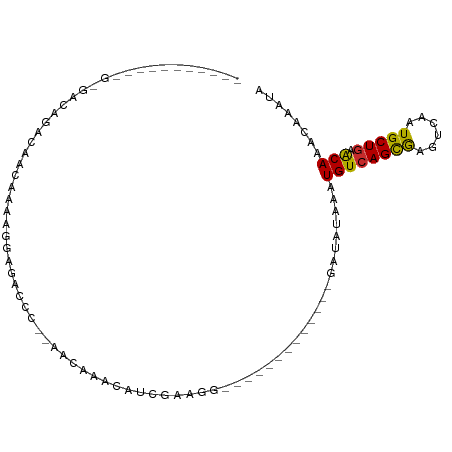

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:32 2011