| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,359,911 – 17,360,007 |
| Length | 96 |
| Max. P | 0.962588 |

| Location | 17,359,911 – 17,360,007 |
|---|---|
| Length | 96 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 73.02 |
| Shannon entropy | 0.55953 |
| G+C content | 0.39563 |
| Mean single sequence MFE | -19.13 |
| Consensus MFE | -10.70 |
| Energy contribution | -10.98 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.962588 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
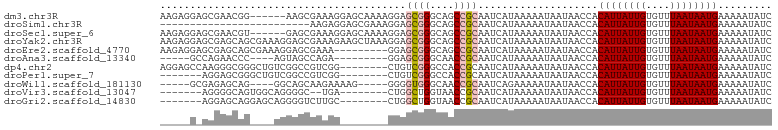
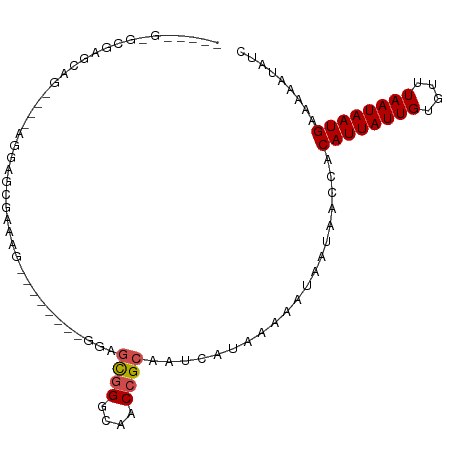
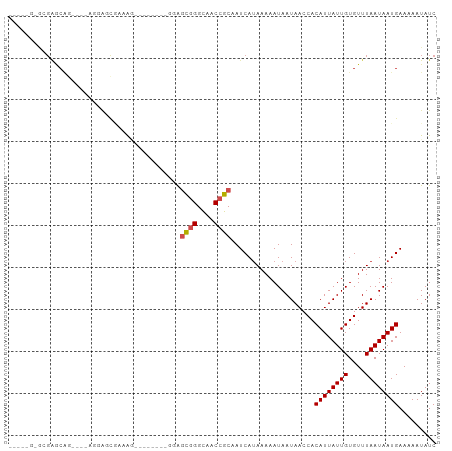
>dm3.chr3R 17359911 96 + 27905053 AAGAGGAGCGAACGG------AAGCGAAAGGAGCAAAAGGAGCGGGCAGCCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC ....((.((...(..------..)(....)..)).......((((....)))).................)).((((((((....))))))))......... ( -18.70, z-score = -2.10, R) >droSim1.chr3R 23408003 77 - 27517382 -------------------------AAGAGGAGCGAAAGGAGCGGGCAGCCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC -------------------------....((..(....)..((((....)))).................)).((((((((....))))))))......... ( -15.60, z-score = -2.52, R) >droSec1.super_6 228752 96 - 4358794 AAGAGGAGCGAACGU------GAGCGAAAGGAGCAAAAGGAGCGGGCAGCCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC ....((.((...(..------..)(....)..)).......((((....)))).................)).((((((((....))))))))......... ( -18.70, z-score = -1.81, R) >droYak2.chr3R 6189357 102 + 28832112 AAGAGGAGCGAGCAGCGAAAGGAGCGAAAGAAGCUAAAGGAGCGGGCAGCCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC ....((....(((..(....)...(....)..)))......((((....)))).................)).((((((((....))))))))......... ( -19.90, z-score = -1.50, R) >droEre2.scaffold_4770 13439856 93 - 17746568 AAGAGGAGCGAGCAGCGAAAGGAGCGAAA---------GGAGCGGGCAGCCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC ....((.....((..(....)..))....---------...((((....)))).................)).((((((((....))))))))......... ( -18.40, z-score = -1.67, R) >droAna3.scaffold_13340 17698119 84 + 23697760 -----GCCAGAACCC----AGUAGCCAGA---------GGAGCGGGCAACCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC -----....((.(((----........).---------)).((((....))))..))................((((((((....))))))))......... ( -16.40, z-score = -2.22, R) >dp4.chr2 3660564 94 + 30794189 AGGAGCCAAGGGCGGGCUGUCGGCCGUCGG--------CUGUCGGGCCACCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC .((.(((...)))((.(((.(((((...))--------))).))).)).))......................((((((((....))))))))......... ( -26.70, z-score = -1.34, R) >droPer1.super_7 3755731 87 - 4445127 -------AGGAGCGGGCUGUCGGCCGUCGG--------CUGUCGGGCCACCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC -------....((((.(((.(((((...))--------))).)))....))))....................((((((((....))))))))......... ( -23.00, z-score = -1.33, R) >droWil1.scaffold_181130 4143230 88 - 16660200 -----GCGAGAGCAG----GGCAGCAAGAAAAG-----GGGGUGGGCAACCGCAAUCAGAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC -----((....)).(----(............(-----(..((((....))))..)).............)).((((((((....))))))))......... ( -19.71, z-score = -2.70, R) >droVir3.scaffold_13047 4664101 85 - 19223366 -------AGGGGCAGUGGCAGGGGC--UGA--------CUGGCUGGUAACCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC -------.(((.(((((((....))--).)--------))).)((((.......))))............)).((((((((....))))))))......... ( -15.90, z-score = 0.10, R) >droGri2.scaffold_14830 5410291 87 + 6267026 -------AGGAGCAGGAGCAGGGGUCUUGC--------CUGGCUGGUAACCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC -------.((.((....)).(.(((..(((--------(.....))))))).).................)).((((((((....))))))))......... ( -17.40, z-score = -0.31, R) >consensus _____G_GCGAGCAG____AGGAGCGAAAG________GGAGCGGGCAACCGCAAUCAUAAAAAUAAUAACCACAUUAUUGUGUUUAAUAAUGAAAAAUAUC .........................................((((....))))....................((((((((....))))))))......... (-10.70 = -10.98 + 0.28)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:29 2011