
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,328,534 – 17,328,614 |
| Length | 80 |
| Max. P | 0.898017 |

| Location | 17,328,534 – 17,328,614 |
|---|---|
| Length | 80 |
| Sequences | 10 |
| Columns | 89 |
| Reading direction | forward |
| Mean pairwise identity | 77.15 |
| Shannon entropy | 0.44714 |
| G+C content | 0.46999 |
| Mean single sequence MFE | -20.63 |
| Consensus MFE | -9.59 |
| Energy contribution | -10.09 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.898017 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17328534 80 + 27905053 AAUAAGCACGAUCAGAGAAGACUUUUAAGGCAACAUGCCUAUCGGCAUGUCAUUGUUG-GCCAACACAUUCGUGCACCAAG-------- .....((((((..((((....))))...(((.(((((((....)))))))((....))-))).......))))))......-------- ( -24.10, z-score = -3.29, R) >droSim1.chr3R 23377166 80 - 27517382 AAUAAGCACGAUCAGAGAAGACUUUUAAGGCAACAUGCCUAUCGGCAUGUCAUUGUUG-GCCAACACAUUCGUGCACCGAG-------- .....((((((..((((....))))...(((.(((((((....)))))))((....))-))).......))))))......-------- ( -24.10, z-score = -3.04, R) >droSec1.super_6 197875 80 - 4358794 AAUAAGCACGAUCAGAGAAGACUUUUAAGGCAACAUGCCUAUCGGCAUGUCAUUGUUG-GCCAACACAUUCGUGCACCGAG-------- .....((((((..((((....))))...(((.(((((((....)))))))((....))-))).......))))))......-------- ( -24.10, z-score = -3.04, R) >droYak2.chr3R 6163711 80 + 28832112 AAUAAGCACGAUCAGAGAAGACUUUUAAGGCAACAUGCCUAUCGGCAUGUCAUUGUUG-GCCAACACAUUCGUGCACCGAG-------- .....((((((..((((....))))...(((.(((((((....)))))))((....))-))).......))))))......-------- ( -24.10, z-score = -3.04, R) >droEre2.scaffold_4770 13408647 80 - 17746568 AAUAAGCACGAUCAGAGAAGACUUUUAAGGCAACAUGCCUAUCGGCAUGUCAUUGUUG-GCCAACACAUUCGUGCACCGCG-------- .....((((((..((((....))))...(((.(((((((....)))))))((....))-))).......))))))......-------- ( -24.10, z-score = -2.68, R) >droAna3.scaffold_13340 17668039 87 + 23697760 AAUAAGCGCGAUCACAGAAGUCUUUUAAGGCAACAAGCCUAUCGGCAUGUCAUUGGAG-GCCAACAU-CCCACAGACCUAUACACAGAG .....((.((((...((....))....((((.....))))))))))..(((..(((.(-(......)-))))..)))............ ( -13.80, z-score = 0.28, R) >dp4.chr2 10768859 63 + 30794189 AAUAAGCGCGAUCAGAGAAGUCUUUUAAGCCAACAUGCCUAUCGGCAUGUCAUUGCAG-ACCCA------------------------- .....((((....((((....))))...))..(((((((....)))))))....))..-.....------------------------- ( -15.00, z-score = -1.71, R) >droPer1.super_0 2116699 63 - 11822988 AAUAAGCGCGAUCAGAGAAGUCUUUUAAGCCAACAUGCCUAUCGGCAUGUCAUUGCAG-GCCCA------------------------- .....((((....((((....))))...))..(((((((....)))))))....))..-.....------------------------- ( -15.00, z-score = -0.87, R) >droVir3.scaffold_12822 3132384 62 - 4096053 AAUAAGCGCGAUCAGAGAAGUCUUUUAAGCCAACAUGCCUGUCGGCAUGUCAUUCGUGCUCA--------------------------- ....(((((((..((((....)))).......(((((((....)))))))...)))))))..--------------------------- ( -20.10, z-score = -2.86, R) >droGri2.scaffold_15074 2843675 84 + 7742996 AAUAAGCGCGAUCAGAGAAGUCUUUUAAGCCAACAUGCCUGUCGGCAUGUCAUUCGCGCACCCAGAC-CUCACACGCACACUAAC---- .....((((((..((((....)))).......(((((((....)))))))...))))))........-.................---- ( -21.90, z-score = -2.46, R) >consensus AAUAAGCACGAUCAGAGAAGACUUUUAAGGCAACAUGCCUAUCGGCAUGUCAUUGUUG_GCCAACAC_UUCGUGCACCGAG________ .......(((((.((((....)))).......(((((((....))))))).)))))................................. ( -9.59 = -10.09 + 0.50)


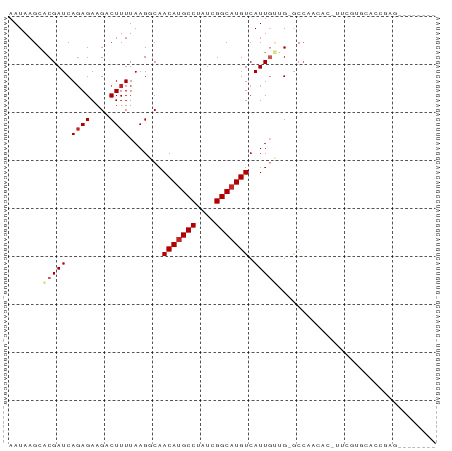
| Location | 17,328,534 – 17,328,614 |
|---|---|
| Length | 80 |
| Sequences | 10 |
| Columns | 89 |
| Reading direction | reverse |
| Mean pairwise identity | 77.15 |
| Shannon entropy | 0.44714 |
| G+C content | 0.46999 |
| Mean single sequence MFE | -22.91 |
| Consensus MFE | -12.08 |
| Energy contribution | -12.18 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.849676 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17328534 80 - 27905053 --------CUUGGUGCACGAAUGUGUUGGC-CAACAAUGACAUGCCGAUAGGCAUGUUGCCUUAAAAGUCUUCUCUGAUCGUGCUUAUU --------......((((((.......(((-.......((((((((....))))))))))).......((......))))))))..... ( -23.91, z-score = -2.12, R) >droSim1.chr3R 23377166 80 + 27517382 --------CUCGGUGCACGAAUGUGUUGGC-CAACAAUGACAUGCCGAUAGGCAUGUUGCCUUAAAAGUCUUCUCUGAUCGUGCUUAUU --------......((((((.......(((-.......((((((((....))))))))))).......((......))))))))..... ( -23.91, z-score = -2.03, R) >droSec1.super_6 197875 80 + 4358794 --------CUCGGUGCACGAAUGUGUUGGC-CAACAAUGACAUGCCGAUAGGCAUGUUGCCUUAAAAGUCUUCUCUGAUCGUGCUUAUU --------......((((((.......(((-.......((((((((....))))))))))).......((......))))))))..... ( -23.91, z-score = -2.03, R) >droYak2.chr3R 6163711 80 - 28832112 --------CUCGGUGCACGAAUGUGUUGGC-CAACAAUGACAUGCCGAUAGGCAUGUUGCCUUAAAAGUCUUCUCUGAUCGUGCUUAUU --------......((((((.......(((-.......((((((((....))))))))))).......((......))))))))..... ( -23.91, z-score = -2.03, R) >droEre2.scaffold_4770 13408647 80 + 17746568 --------CGCGGUGCACGAAUGUGUUGGC-CAACAAUGACAUGCCGAUAGGCAUGUUGCCUUAAAAGUCUUCUCUGAUCGUGCUUAUU --------......((((((.......(((-.......((((((((....))))))))))).......((......))))))))..... ( -23.91, z-score = -1.48, R) >droAna3.scaffold_13340 17668039 87 - 23697760 CUCUGUGUAUAGGUCUGUGGG-AUGUUGGC-CUCCAAUGACAUGCCGAUAGGCUUGUUGCCUUAAAAGACUUCUGUGAUCGCGCUUAUU ....((((((((((((..(((-..(((((.-..)))))((((.(((....))).)))).)))....))))..)))))..)))....... ( -21.80, z-score = -0.45, R) >dp4.chr2 10768859 63 - 30794189 -------------------------UGGGU-CUGCAAUGACAUGCCGAUAGGCAUGUUGGCUUAAAAGACUUCUCUGAUCGCGCUUAUU -------------------------.((((-((((...((((((((....)))))))).)).....))))))................. ( -17.80, z-score = -1.51, R) >droPer1.super_0 2116699 63 + 11822988 -------------------------UGGGC-CUGCAAUGACAUGCCGAUAGGCAUGUUGGCUUAAAAGACUUCUCUGAUCGCGCUUAUU -------------------------(((((-(......((((((((....))))))))))))))......................... ( -17.80, z-score = -1.23, R) >droVir3.scaffold_12822 3132384 62 + 4096053 ---------------------------UGAGCACGAAUGACAUGCCGACAGGCAUGUUGGCUUAAAAGACUUCUCUGAUCGCGCUUAUU ---------------------------(((((.(((..((((((((....))))))))........(((....)))..))).))))).. ( -21.20, z-score = -3.34, R) >droGri2.scaffold_15074 2843675 84 - 7742996 ----GUUAGUGUGCGUGUGAG-GUCUGGGUGCGCGAAUGACAUGCCGACAGGCAUGUUGGCUUAAAAGACUUCUCUGAUCGCGCUUAUU ----......(((((.(.(((-((((...((.((...(((((((((....))))))))))).))..))))))).)....)))))..... ( -31.00, z-score = -2.26, R) >consensus ________CUCGGUGCACGAA_GUGUUGGC_CAACAAUGACAUGCCGAUAGGCAUGUUGCCUUAAAAGACUUCUCUGAUCGCGCUUAUU ......................................((((((((....))))))))............................... (-12.08 = -12.18 + 0.10)


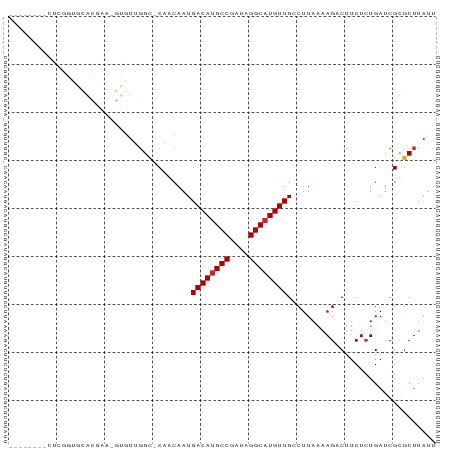
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:21 2011