
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,298,499 – 17,298,589 |
| Length | 90 |
| Max. P | 0.872706 |

| Location | 17,298,499 – 17,298,589 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 71.20 |
| Shannon entropy | 0.52955 |
| G+C content | 0.58748 |
| Mean single sequence MFE | -34.58 |
| Consensus MFE | -18.17 |
| Energy contribution | -18.15 |
| Covariance contribution | -0.02 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.872706 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 17298499 90 - 27905053 GCAGACGUG---------GGCGUGACUUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAA-UCGCAUACAAGU--------------AUUCGUAUUCGGAUUUGGACUCGGA .......((---------(((((((.((((.((((((..(.((...)).)..))))))))))-))))...(((..--------------(((((....))))))))).)))).. ( -30.90, z-score = -1.28, R) >droSim1.chr3R 23347293 100 + 27517382 GCAGACGGG---------GGCGUGACUUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAA-UCGCAUACGGAUUCGUAUG----AGUAUUCGUAUUCGGAUUUGGACUCGGA .....((((---------...((((.((((.((((((..(.((...)).)..))))))))))-))))...((((((((((((----......))))..))))))))..)))).. ( -39.30, z-score = -2.80, R) >droSec1.super_6 168355 104 + 4358794 GCAGACGGG---------GGCGUGACUUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAA-UCGCAUACGGAUUCCUACGAGUAUUCAUUCGUAUUCGGAUUUGGACUCGGA .....((((---------...((((.((((.((((((..(.((...)).)..))))))))))-))))...(((((((.(((((((....)))))))...)))))))..)))).. ( -41.00, z-score = -2.89, R) >droYak2.chr3R 6133647 94 - 28832112 GCAGACGGG---------GGCGUGACUUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAA-UCGCAUUCGGAUU----------UGGAUUCGUAAUCGGAUGUGGACUCGGA ((.......---------.))((((.((((.((((((..(.((...)).)..))))))))))-)))).((((..(.----------.(.(((((....))))).)..)..)))) ( -36.50, z-score = -2.03, R) >droEre2.scaffold_4770 13378125 95 + 17746568 GCAGACGGG---------GGCGUGACUUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAAUUCGUAUUCGCAUU----------GGGAUUCGUAUUCGGAUUUGGACUCGCA ...(.((((---------.((((((.((((.((((((..(.((...)).)..)))))))))).)))....))).((----------.(((((((....))))))).))))))). ( -36.10, z-score = -2.26, R) >droAna3.scaffold_13340 17641406 88 - 23697760 GCAGACGGG---------GGCGUGACUUCCACAGGCUUUAUGCUUGGCUGGGGGCCUGGGAA------UACGGAC-----------CGGACCCGGACCAGGGCCCGAAAUCGGA .(((.((((---------((.....))))).(((((.....)))))))))(((.(((((...------..(((..-----------.....)))..))))).)))......... ( -34.70, z-score = -0.31, R) >dp4.chr2 10739953 80 - 30794189 CCAGACGGG---------GGCGUGACAUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAA------------------------UCGCCCUGUAUCCGAAC-CGAUCUCGCA ..(((((((---------((((.....(((.((((((..(.((...)).)..))))))))).------------------------.)))))((....))..)-)).))).... ( -31.80, z-score = -0.63, R) >droWil1.scaffold_181089 8168452 85 - 12369635 GAAGGCGGGUCCCAACCUCGUAUCUCAUCCACAGGCUUUAUGCUUGGCUGGGGGCUUGGGAAUC----------------------GGCAUCGGCAUCGGAAUCAGA------- ((.(.((..(((((((((((.....).....(((((.....)))))....)))).))))))..)----------------------).).))...............------- ( -26.30, z-score = 0.34, R) >consensus GCAGACGGG_________GGCGUGACUUCCACAGGCUUUCUGCUUGGCUGGGGGCCUGGGAA_UCGCAUACGGAU___________UGGAUUCGUAUUCGGAUUUGGACUCGGA ....(((((.........(......).(((.(((((((((.((...)).)))))))))))).............................)))))................... (-18.17 = -18.15 + -0.02)


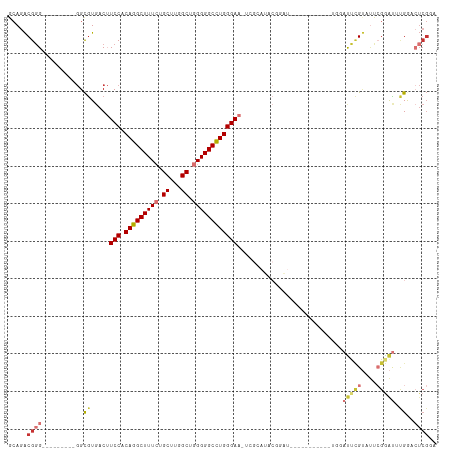
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:16 2011