| Sequence ID | dm3.chr3R |
|---|---|
| Location | 17,200,482 – 17,200,579 |
| Length | 97 |
| Max. P | 0.993158 |

| Location | 17,200,482 – 17,200,579 |
|---|---|
| Length | 97 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 58.47 |
| Shannon entropy | 0.85474 |
| G+C content | 0.37719 |
| Mean single sequence MFE | -21.39 |
| Consensus MFE | -11.77 |
| Energy contribution | -11.20 |
| Covariance contribution | -0.57 |
| Combinations/Pair | 1.91 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.993158 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
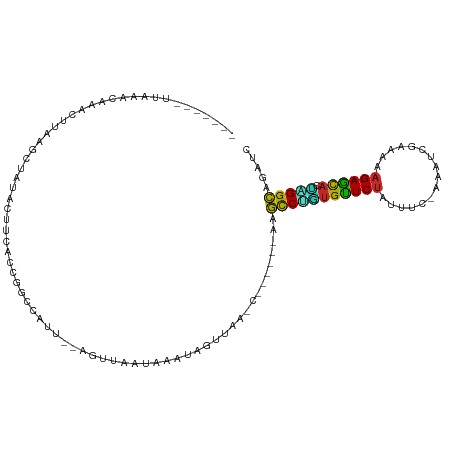
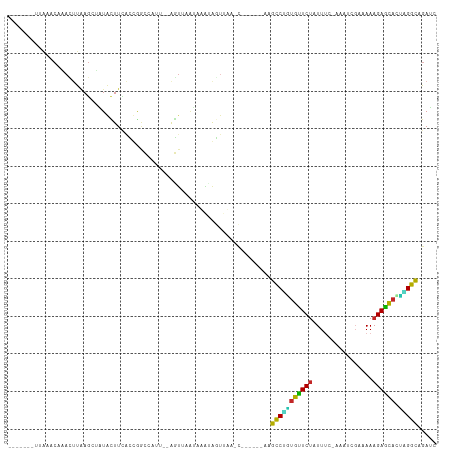
>dm3.chr3R 17200482 97 - 27905053 -------UUGAACAAACUUAAGCUAUACUUCGCUGGCCAUU--AGUUAAUAAGUAGUUUA-C------GGGCCUGUGUUCUAUUUC-GAAUCGACAAAGAGCACUAGGCAGAUC -------.........(.((((((((..((.((((.....)--))).))...))))))))-.------).(((((((((((.(.((-(...))).).))))))).))))..... ( -22.70, z-score = -1.06, R) >droSim1.chr3R 23243055 97 + 27517382 -------UUAAACAAACUUAAGCUGUACUUCGCUGGCAAUU--AGUUAAUGAAUAGUUUA-C------GGGCCUGUGUUCUAUUUC-GAAGCGACAAAGAGCACUAGGCAGAUC -------.........(.((((((((...(((.((((....--.)))).)))))))))))-.------).(((((((((((.(.((-(...))).).))))))).))))..... ( -24.70, z-score = -1.47, R) >droSec1.super_6 68602 97 + 4358794 -------UUAAACAAACUUAAGCUGUACUUCGCUGGCAAUU--AGUUAAUGAAUAGUUUA-C------GGGCCUGUGUUCUAUUUC-GAAGCGACAAAGAGCACUAGGCAGAUC -------.........(.((((((((...(((.((((....--.)))).)))))))))))-.------).(((((((((((.(.((-(...))).).))))))).))))..... ( -24.70, z-score = -1.47, R) >droYak2.chr3R 6006391 97 - 28832112 -------UUAAACAAAGUUCAGCUAUACUUCACCGACUCGU--AGUAAAUAAGCAGUUUA-C------CAGCCUGUGUUCUAUUUG-AAAUCGAUAAAGAGCACUAGGCAGAUU -------..............(((.((((.(........).--))))....)))......-.------..(((((((((((..(((-....)))...))))))).))))..... ( -20.20, z-score = -1.58, R) >droEre2.scaffold_4770 13275604 97 + 17746568 -------UAAUACAAACUUAAGCUAUACUUCACUGACUUAU--AGUUAAUAAGUAGUUAA-C------AAGCCUGUGUUCUAUUUG-AAAUCGAUAAAGAGCACUAGGCAGAUC -------........((((((((((((..((...))..)))--))))..)))))......-.------..(((((((((((..(((-....)))...))))))).))))..... ( -23.70, z-score = -2.80, R) >droAna3.scaffold_13340 216216 96 + 23697760 ------UGCAAUCAAAUGUAAGCAAUACUU--CUGGCCAUU--AGUAUUUAAUUAGUUAAGC------CAGCCUGUGUUCUAUUUU-AAA-CGCUAAAGAACAUCUGGCCGAUC ------((((......))))..........--.((((.(((--(((.....))))))...))------))(((.(((((((.....-...-......)))))))..)))..... ( -17.14, z-score = -0.31, R) >dp4.chr2 5698264 86 - 30794189 ------------------CGAUUCUUAUUUAAGCGUUGCCU--AACAAGCCAGUAUUUAAGC------AAGCCUGUGUUCUAUUAA-ACUCCGCAAAAGAACA-UGGGCCGAUC ------------------.((((....((((((..(((.((--....)).)))..)))))).------..(((((((((((.....-..........))))))-))))).)))) ( -18.36, z-score = -1.10, R) >droVir3.scaffold_12822 2983350 106 + 4096053 UUGUUGCCCGUGCACAUUUAAGAGCUGCUUUAUCAACCAUUUAGAU--AGAUAAGAUAAAAA---AAAAAGCCAAUGCUCUAUUUU---CGCUUAAAAGAGCAUUUGGCCGAGC .(((.((....))))).......(((..((((((.........)))--)))...........---.....(((((((((((.(((.---.....))))))))).)))))..))) ( -19.60, z-score = -0.08, R) >droMoj3.scaffold_6540 393954 104 + 34148556 ------UCAAUGCAUAUUUAAGAGCUGUUUUAUCAGCCAUUUUAGUUAAGAUAAGAUUAGCUUCUGGUAGAUCGCCGCUCAAUUU----UUCCGAAAAGAGUUUGCGGUCGAGC ------.....((.((((...((((((((((((((((.......)))..))))))).))))))..))))..(((((((..(((((----((.....))))))).)))).))))) ( -23.40, z-score = -0.71, R) >droGri2.scaffold_15074 2713445 112 - 7742996 -UAAUUGCAAUGCACAUUUAAGAGCUGCUUUAUCAGCCAUUUUAGUUUAAAUAAGUACAACUUUUAA-AAGCCUUUGCUCUAUUCUCGAUUCGCAAAAGAGUAUUUGGCUGAGG -.....(((..((.(......).)))))....(((((((.....((((....(((.....)))....-))))...((((((................))))))..))))))).. ( -19.39, z-score = 0.68, R) >consensus _______UUAAACAAACUUAAGCUAUACUUCACCGGCCAUU__AGUUAAUAAAUAGUUAA_C______AAGCCUGUGUUCUAUUUC_AAAUCGAAAAAGAGCACUAGGCAGAUC ......................................................................(((((((((((................))))))).))))..... (-11.77 = -11.20 + -0.57)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:32:06 2011