
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 16,983,510 – 16,983,612 |
| Length | 102 |
| Max. P | 0.639610 |

| Location | 16,983,510 – 16,983,612 |
|---|---|
| Length | 102 |
| Sequences | 7 |
| Columns | 122 |
| Reading direction | reverse |
| Mean pairwise identity | 59.64 |
| Shannon entropy | 0.76841 |
| G+C content | 0.44631 |
| Mean single sequence MFE | -23.42 |
| Consensus MFE | -10.05 |
| Energy contribution | -10.10 |
| Covariance contribution | 0.05 |
| Combinations/Pair | 1.86 |
| Mean z-score | -0.62 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.639610 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
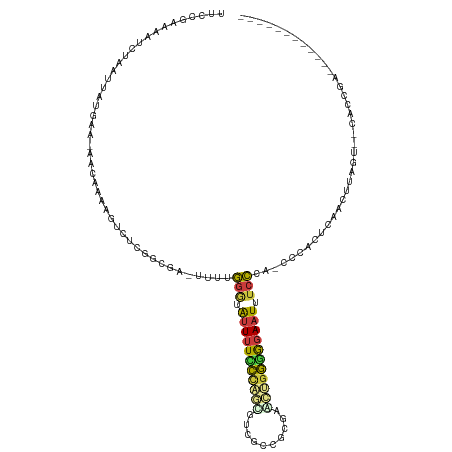
>dm3.chr3R 16983510 102 - 27905053 UUCUGAAAAUCUAAUUAUGAA----AAAAUUUUGGGCGA-UUUUGGGUAUUUUCCCAGCGUCGCCGCGAACUGGGGGAAUUUCCCA-CCCACUCAACUUAGU--CACCGA------------ ..........((((...(((.----.....(((((((((-(.(((((......))))).)))))).)))).((((((......)).-)))).)))..)))).--......------------ ( -29.20, z-score = -1.67, R) >droAna3.scaffold_12911 2232699 106 - 5364042 ------------CCCCACAAAUCACAAGAAUGUCCCCCU-UUUCGGGUGUUUUUCCAACGAC-CAACUGACAGGGGGAAUUUCCCA-CCCACUCG-CCCAGUCCCGCCCACUGAGUCACAAC ------------...............(((..(((((((-..((((..(((........)))-...)))).)))))))..)))...-.......(-(.((((.......)))).))...... ( -21.50, z-score = 0.60, R) >droEre2.scaffold_4770 13069220 105 + 17746568 UGCCCAAAAUCAAAUUAUAAA-AACCGAAGUGUCCGCGA-UUUUGGGUAUUUUCCCAGUGUCGCCGCGAAAAGGGGGAAUUUCCCA-CCCACUCGACUUAGU--CACCAA------------ .....................-...(((.(((...((((-(.(((((......))))).)))))........(((((.....))).-)))))))).......--......------------ ( -26.40, z-score = -1.08, R) >droYak2.chr3R 5766367 103 - 28832112 UUUCAAAAAUCUAACUAUGAA-AUCCAAAGUGUCCGCGA-UUUUGGGUAUUUU-CCAGCGUCGCCGCGAACU-GGGGAAUUUCCCA-CCCACUCAACUUAGU--CACCGA------------ ..................(((-((((..(((..(.((((-(.(((((.....)-)))).))))).)...)))-..)).)))))...-...............--......------------ ( -20.90, z-score = -0.04, R) >droSec1.super_38 252462 107 + 400794 UUCCGAAAAUCUUAUUAUGAA-GACAAAAGUCUCGGCGAUUUUUGAAUAUUUUCCCAGCGUAGCCGCCAGCUGGGGGAAUUUCCCACCCCACUCAACUUAGU--CACCGA------------ .....................-(((....)))((((.((((.((((..(((..((((((..........))))))..)))............))))...)))--).))))------------ ( -25.24, z-score = -1.14, R) >droSim1.chr3R 23038266 106 + 27517382 UUCCGAAAAUCUAAUUAUGAA-GACAAAAGUCUCGGCGAUUUUUGGGUAUUUUCCCAGCGUCGCCGCCAGCUGGGGGAAUUUCCCA-CCCACUCAACUUAGU--CACCGA------------ .....................-(((...(((..(((((((..(((((......))))).)))))))...)))(((((......)).-)))..........))--).....------------ ( -30.10, z-score = -1.58, R) >droVir3.scaffold_13047 14760635 87 + 19223366 -UUCGAAAAAGUAUCUAUAUAAUAAGAAUAUCGAGUUUGCUUUUUGCCGAUCUUUUGUAAGAUUUAUAAUGCGAAAAAGUGACAUAAA---------------------------------- -((((((((((((((.((((.......)))).))...))))))))...((((((....)))))).......)))).............---------------------------------- ( -10.60, z-score = 0.57, R) >consensus UUCCGAAAAUCUAAUUAUGAA_AACAAAAGUCUCGGCGA_UUUUGGGUAUUUUCCCAGCGUCGCCGCGAACUGGGGGAAUUUCCCA_CCCACUCAACUUAGU__CACCGA____________ ............................................(((.(((((((((((..........))))))))))).)))...................................... (-10.05 = -10.10 + 0.05)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:31:33 2011