| Sequence ID | dm3.chr3R |
|---|---|
| Location | 16,817,972 – 16,818,063 |
| Length | 91 |
| Max. P | 0.922082 |

| Location | 16,817,972 – 16,818,063 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 64.01 |
| Shannon entropy | 0.66082 |
| G+C content | 0.53007 |
| Mean single sequence MFE | -23.61 |
| Consensus MFE | -12.91 |
| Energy contribution | -13.52 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.65 |
| Mean z-score | -0.93 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.922082 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
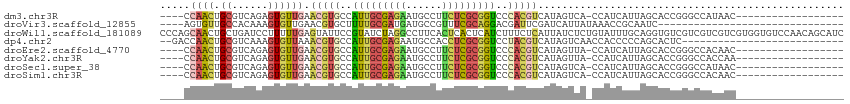
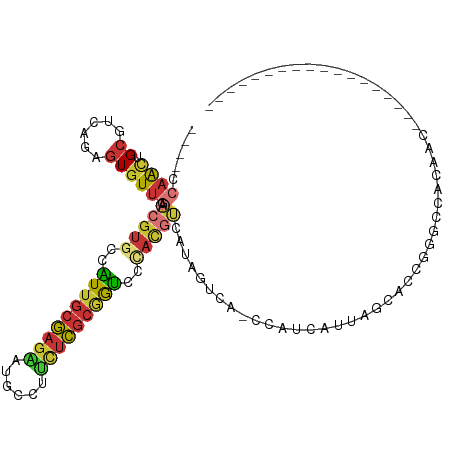
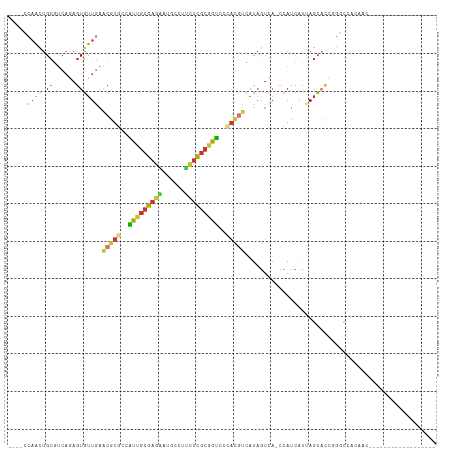
>dm3.chr3R 16817972 91 + 27905053 ----CCAACUGCGUCAGAGUGUUGAACGUGCCAUUGCGAGAAUGCCUUCUCGCGGUCCCACGUCAUAGUCA-CCAUCAUUAGCACCGGGCCAUAAC------------------ ----.....((.(((.(.((((((((((((..((((((((((....))))))))))..)))))........-......)))))))).)))))....------------------ ( -26.93, z-score = -1.53, R) >droVir3.scaffold_12855 2312137 79 + 10161210 ----AGUGUUGCCACAAAGUGUUGAACGUGCUUUUGCGAUGAUGCCGUUUCGCAGGACGAUUCGAUCAUUAUAAACCGCAAUC------------------------------- ----...(((((.....((((((((((((.((...((((..........)))))).))).))))).)))).......))))).------------------------------- ( -16.90, z-score = 0.62, R) >droWil1.scaffold_181089 6145396 114 + 12369635 CCCAGCAACUGCUGAUCCUUUUUGAGUAUUCCGUAUCUAGGCCUUCACUCACUCAUCUUUCUCAUUAUCUCUGUAUUUGCAGGUGUCGUCGUCGUCGUGGUGUCCAACAGCAUC .........(((((........(((((...((.......)).....)))))...........((((((.(((((....))))).(.((....)).))))))).....))))).. ( -17.90, z-score = 0.79, R) >dp4.chr2 18173997 85 - 30794189 --GACCAACUGCGUCAAAGUGUUAAACGUGCCAUUGCGAGAAUGCCACCUCGCGGUCCUACGUCAUAGUCAACCACCCCCAGCACUC--------------------------- --((((....).)))..((((((..(((((..((((((((........))))))))..)))))....((.....))....)))))).--------------------------- ( -19.70, z-score = -1.52, R) >droEre2.scaffold_4770 12907744 91 - 17746568 ----CCAACUGCGUCAGAGUGUUGAACGUGCCAUUGCGAGAAUGCCUUCUCGCGGUCCCACGUCAUAGUUA-CCAUCAUUAGCACCGGGCCACAAC------------------ ----.....((.(((.(.((((((((((((..((((((((((....))))))))))..)))))........-......)))))))).)))))....------------------ ( -26.83, z-score = -1.47, R) >droYak2.chr3R 5610036 91 + 28832112 ----CCAACUGCGUCAGAGUGUUGAACGUGCCAUUGCGAGAAUGCCUUCUCGCGGUCCCACGUCAUAGUUA-CCAUCAUUAGCACCGGGCCACCAA------------------ ----.....((.(((.(.((((((((((((..((((((((((....))))))))))..)))))........-......)))))))).)))))....------------------ ( -26.83, z-score = -1.51, R) >droSec1.super_38 92858 91 - 400794 ----CCAACUGCGUCAGAGUGUUGAACGUGCCAUUGCGAGAAUGCCUUCUCGCGGUCCCACGUCAUAGUCA-CCAUCAUUAGCACCGGGCCAUAAC------------------ ----.....((.(((.(.((((((((((((..((((((((((....))))))))))..)))))........-......)))))))).)))))....------------------ ( -26.93, z-score = -1.53, R) >droSim1.chr3R 22876452 91 - 27517382 ----CCAACUGCGUCAGAGUGUUGAACGUGCCAUUGCGAGAAUGCCUUCUCGCGGUCCCACGUCAUAGUCA-CCAUCAUUAGCACCGGGCCACAAC------------------ ----.....((.(((.(.((((((((((((..((((((((((....))))))))))..)))))........-......)))))))).)))))....------------------ ( -26.83, z-score = -1.31, R) >consensus ____CCAACUGCGUCAGAGUGUUGAACGUGCCAUUGCGAGAAUGCCUUCUCGCGGUCCCACGUCAUAGUCA_CCAUCAUUAGCACCGGGCCACAAC__________________ .....((((.((......)))))).(((((..(((((((((......)))))))))..)))))................................................... (-12.91 = -13.52 + 0.61)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:31:22 2011