
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 16,631,668 – 16,631,725 |
| Length | 57 |
| Max. P | 0.500000 |

| Location | 16,631,668 – 16,631,725 |
|---|---|
| Length | 57 |
| Sequences | 14 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 91.79 |
| Shannon entropy | 0.20737 |
| G+C content | 0.47559 |
| Mean single sequence MFE | -10.58 |
| Consensus MFE | -9.33 |
| Energy contribution | -9.05 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.59 |
| Mean z-score | -1.09 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
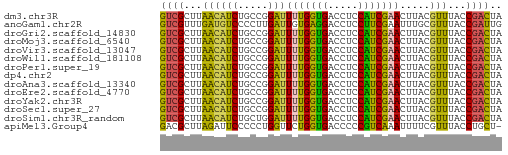
>dm3.chr3R 16631668 57 - 27905053 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >anoGam1.chr2R 4729700 57 - 62725911 GUCGUUUGAUGUCCCCUUGAUUGUGAGGACCUCCUUCGAAUUUGCGUUUACCGAUUG .(((...((.((((.(........).)))).))...))).................. ( -9.30, z-score = -0.77, R) >droGri2.scaffold_14830 1131985 57 - 6267026 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droMoj3.scaffold_6540 29702742 57 - 34148556 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droVir3.scaffold_13047 7145261 57 + 19223366 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droWil1.scaffold_181108 2474795 57 + 4707319 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droPer1.super_19 1706559 57 - 1869541 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >dp4.chr2 5970540 57 - 30794189 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droAna3.scaffold_13340 10746350 57 + 23697760 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droEre2.scaffold_4770 12714091 57 + 17746568 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droYak2.chr3R 5426781 57 - 28832112 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droSec1.super_27 695129 57 + 799419 GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.24, R) >droSim1.chr3R_random 823497 57 - 1307089 GUCGCUUAACAUCUGCUGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -11.00, z-score = -1.45, R) >apiMel3.Group4 9923366 56 + 10796202 GACGCUUAGAUUCCCCCUGGUUCUGGUGACCCCCGUCAAAUUUUCGUUUACCUGCU- ((((...(((..((....)).)))(....)...))))...................- ( -6.80, z-score = 0.50, R) >consensus GUCGCUUAACAUCUGCCGGAUUUUGGUGACCUCCAUCGAACUUACGUUUACCGACUA ((((...((((((.....)))(((((((.....))))))).....)))...)))).. ( -9.33 = -9.05 + -0.28)

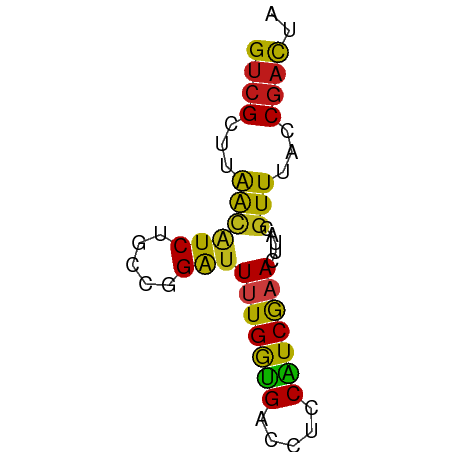
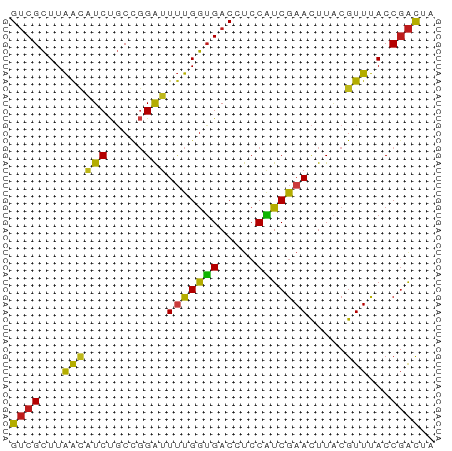
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:30:53 2011