
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 16,573,079 – 16,573,162 |
| Length | 83 |
| Max. P | 0.523532 |

| Location | 16,573,079 – 16,573,162 |
|---|---|
| Length | 83 |
| Sequences | 15 |
| Columns | 84 |
| Reading direction | reverse |
| Mean pairwise identity | 81.69 |
| Shannon entropy | 0.44181 |
| G+C content | 0.52912 |
| Mean single sequence MFE | -25.16 |
| Consensus MFE | -14.28 |
| Energy contribution | -13.82 |
| Covariance contribution | -0.46 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.523532 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 16573079 83 - 27905053 UACAUGUACUUAAUGCAGUGGGUGGCCCCCACCCACGACGCGGUCAUCAGGAUGGUCACGCACACUCCGAAGUACAUCUGUAA- (((((((((((......((((((((...))))))))(.((.(..(((....)))..).)))........)))))))..)))).- ( -30.90, z-score = -2.42, R) >apiMel3.GroupUn 324543927 83 + 399230636 UAUAAAUAUUUAAUACAAUGUGUUGCACCGACCCAACUUGCUGUAACACAUAUUGUGACGCACACUCCAAAAUAUAUCUGAAA- .....((((((......((((((((((.(((......))).))))))))))...(((.....)))....))))))........- ( -15.80, z-score = -2.79, R) >droGri2.scaffold_14830 1064010 81 - 6267026 UACAUGUACUUGAUGCAGUGCGUCGCACCCACCCAAGAGGCAGUCACCAGAAUCGUAACGCAUACGCCAAAGUACAUCUGG--- ...((((((((...((.((((((..(......((....))...((....))...)..))))))..))..))))))))....--- ( -19.40, z-score = -0.88, R) >droMoj3.scaffold_6540 29647437 84 - 34148556 UACAUGUACUUGAUGCAGUGCGUGGCGCCCACCCACGAAGCAGUCACUAGAAUCGUGACGCACACCCCAAAGUACAUCUGUAGC (((((((((((......((((((((.(....)))))......(((((.......)))))))))......)))))))..)))).. ( -26.60, z-score = -2.38, R) >droVir3.scaffold_13047 7090234 83 + 19223366 UACAUGUACUUGAUGCAGUGCGUGGCGCCCACCCAUGAGGCAGUCACUAGAAUGGUAACGCACACGCCAAAGUACAUCUGCCC- ...((((((((...((.((((((((((((((....)).))).)))(((.....))).))))))..))..))))))))......- ( -26.10, z-score = -1.14, R) >droWil1.scaffold_181108 2413792 83 + 4707319 UACAUAUACUUGAUGGUAUGAGUGGCUCCUACCCAUGACCCGGUCACCAGAAUGGUCACGCACACACCAUAGUACAUCUGCAG- ......((((..(((((.((.((((.......))))((((...((....))..)))).....)).))))))))).........- ( -16.30, z-score = 0.99, R) >droPer1.super_19 1647829 83 - 1869541 UACAUGUACUUGAUGCAGUGCGUGGCCCCCACCCAUGAGGCGGUCAUCAGGAUGGUCACGCACACCCCGAAGUACAUCUGCAA- ...((((((((......((((((((((.((.((.....)).))((.....)).))))))))))......))))))))......- ( -28.10, z-score = -0.99, R) >dp4.chr2 5910637 83 - 30794189 UACAUGUACUUGAUGCAGUGCGUGGCCCCCACCCACGAGGCGGUCAUCAGGAUGGUCACGCACACCCCGAAGUACAUCUGCAA- ...((((((((......((((((((((.((.((.....)).))((.....)).))))))))))......))))))))......- ( -28.10, z-score = -1.08, R) >droAna3.scaffold_13340 10686785 83 + 23697760 UACAUGUACUUGAUGCAAUGCGUGGCCCCCACCCACGAGGCAGUCAUCAGGAUGGUCACGCACACCCCGAAGUACAUCUGAAG- ...((((((((.......(((((((((.((....((......)).....))..))))))))).......))))))))......- ( -23.54, z-score = -0.99, R) >droEre2.scaffold_4770 12656329 83 + 17746568 UACAUGUACUUAAUGCAGUGGGUGGCCCCCACCCACGAGGCGGUCAUCAGGAUGGUCACGCACACUCCGAAGUACAUCUGUGA- (((((((((((..(((.((((((((...))))))))..(((.(((.....))).)))..))).......)))))))..)))).- ( -31.50, z-score = -1.82, R) >droYak2.chr3R 5365545 83 - 28832112 UACAUGUACUUAAUGCAGUGGGUGGCCCCCACCCACGAGGCGGUCAUCAGGAUGGUCACGCACACUCCGAAGUACAUCUGUGA- (((((((((((..(((.((((((((...))))))))..(((.(((.....))).)))..))).......)))))))..)))).- ( -31.50, z-score = -1.82, R) >droSec1.super_27 636352 83 + 799419 UACAUGUACUUAAUGCAGUGGGUGGCCCCCACCCACGAGGCGGUCAUCAGGAUGGUCACGCACACUCCGAAGUACAUCUGUGC- (((((((((((..(((.((((((((...))))))))..(((.(((.....))).)))..))).......)))))))..)))).- ( -31.20, z-score = -1.69, R) >droSim1.chr3R 22626827 83 + 27517382 UACAUGUACUUAAUGCAGUGGGUGGCCCCCACCCACGAGGCGGUCAUCAGGAUGGUCACGCACACUCCGAAGUACAUCUGUGA- (((((((((((..(((.((((((((...))))))))..(((.(((.....))).)))..))).......)))))))..)))).- ( -31.50, z-score = -1.82, R) >triCas2.ChLG1X 16886 79 + 8109244 UACAAGUAUUUUAUGCAGUGGGUGGCCCCCACCCAAGAGGCUGUCACGCAGAUUGUCACCCAUAUCCCGAAGUAAAUCU----- ......((((((.....(((((((((......((....))(((.....)))...))))))))).....)))))).....----- ( -18.70, z-score = -0.86, R) >anoGam1.chr2R 3436366 77 + 62725911 UACAGGAACUUGAUGCAGUGCGUUGCACCAACCCACGAGGCCGUUACCAGUACGGUUACGCACACGCCGAAGUACAU------- ....((....((.(((.(((.((((...)))).)))(..(((((.......)))))..))))))..)).........------- ( -18.20, z-score = 0.16, R) >consensus UACAUGUACUUGAUGCAGUGCGUGGCCCCCACCCACGAGGCGGUCAUCAGGAUGGUCACGCACACUCCGAAGUACAUCUGUAA_ .....((((((..(((.(((.((((...)))).)))...)))((.(((.....))).))..........))))))......... (-14.28 = -13.82 + -0.46)


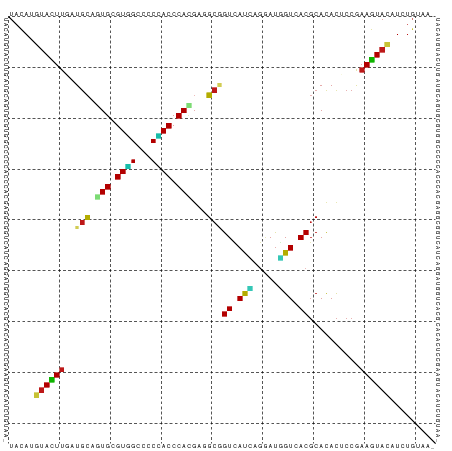
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:30:41 2011