| Sequence ID | dm3.chr3R |
|---|---|
| Location | 16,405,565 – 16,405,701 |
| Length | 136 |
| Max. P | 0.962285 |

| Location | 16,405,565 – 16,405,701 |
|---|---|
| Length | 136 |
| Sequences | 5 |
| Columns | 136 |
| Reading direction | forward |
| Mean pairwise identity | 85.29 |
| Shannon entropy | 0.26264 |
| G+C content | 0.46369 |
| Mean single sequence MFE | -42.22 |
| Consensus MFE | -31.56 |
| Energy contribution | -32.52 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.959363 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
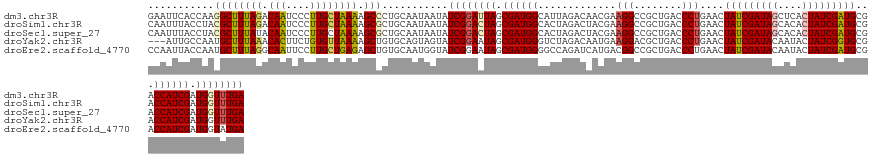
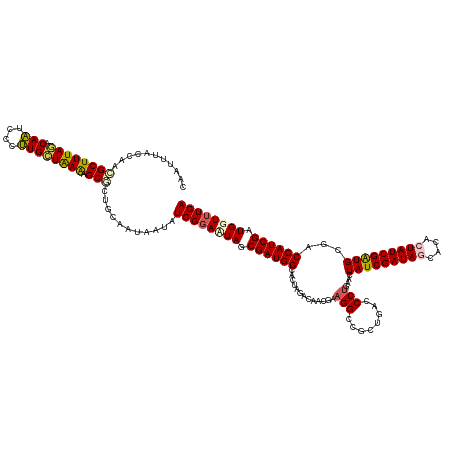
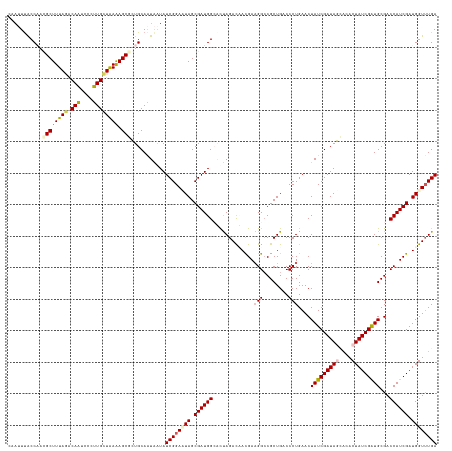
>dm3.chr3R 16405565 136 + 27905053 UCAAACCAUCGAUGGUCGCAUCGAUAGUGAGCUAUCGAUAGUUCAGGGUCAGCGGCCUUCGUUGUCUAAUGCCAUCGCUAAUCCGAUAUUAUUGCAGGGCUUUUAGCAAGGGAUUGUCUAAAGCCUUGGUGAAUUC (((...((.(((.((((((...(((..(((((((....)))))))..))).)))))).))).))..(((((..((((......))))..)))))(((((((((..((((....))))..))))))))).))).... ( -44.10, z-score = -2.24, R) >droSim1.chr3R 22454056 136 - 27517382 UCAAACCAUCGAUGGUCGCAUCGAUAGUGUGCUAUCGAUAGUUCAGGGUCAGCGGCCUUCGUAGUCUAGUGCCAUCGCUAGUCCGAUAUUAUUGCAGCGCUUUUAGCAAGGGAUUGUCUAAAGCGUAGGUAAAUUG ....((((((((.((((((((((((((....)))))))).(........).)))))).))...(.((((((....)))))).).))).........(((((((..((((....))))..))))))).)))...... ( -44.30, z-score = -2.63, R) >droSec1.super_27 468480 136 - 799419 UCAAACCAUCGAUGGUCGCAUCGAUAGUGUGCUAUCGAUAGUUCAGGGUCAGCGGCCUUCGUAGUCUAGUGCCAUCGCUAGUCCGAUAUUAUUGCAGCGCUUUUAGCAAGGGAUUGUAUAAAGCGUAGGUAAAUUG ....((((((((.((((((((((((((....)))))))).(........).)))))).))...(.((((((....)))))).).))).........(((((((..((((....))))..))))))).)))...... ( -42.90, z-score = -2.28, R) >droYak2.chr3R 5191648 133 + 28832112 UCAAACCAUCGAUGGUCGCACCGAUAGUAUUGUAUCGAUAGUUCAGGGUCAGCGUCCUUCAUUGUCUAGACCCAUCGCUAUUCCGAUACUACUGCACAGCUUUUAACACAGAAGUGUUUAAAGCAUUGGCAAU--- ....((((....))))....(((((.(((..(((((((((((...(((((((((........)).)).)))))...)))))..))))))...)))...(((((.(((((....))))).))))))))))....--- ( -38.40, z-score = -3.29, R) >droEre2.scaffold_4770 12479140 136 - 17746568 UCAUACCAUCGAUGGUCGCAUCGAUAGUAUUGUAUCGAUAGUUCAGGGUCAGCGGCCGUCAUGAUCUGGCCCCAUCGCUAUUCCGAUACCAUUGCACAGCUCUCAGCAAGGAAUUGCCUAAAGCAUUGGUAAUUGG ......(((.((((((((((((((((......))))))).(........).))))))))))))...((((......))))..(((((((((.(((...((.....)).(((.....)))...))).)))).))))) ( -41.40, z-score = -1.50, R) >consensus UCAAACCAUCGAUGGUCGCAUCGAUAGUGUGCUAUCGAUAGUUCAGGGUCAGCGGCCUUCGUAGUCUAGUGCCAUCGCUAGUCCGAUAUUAUUGCAGAGCUUUUAGCAAGGGAUUGUCUAAAGCAUUGGUAAAUUG ....((.(((((((....))))))).)).......(((((((...(((..((((.....................))))..)))...)))))))(((((((((..((((....))))..)))))))))........ (-31.56 = -32.52 + 0.96)
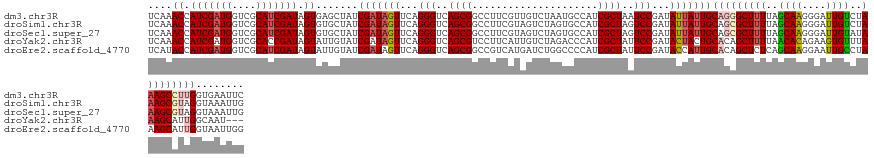
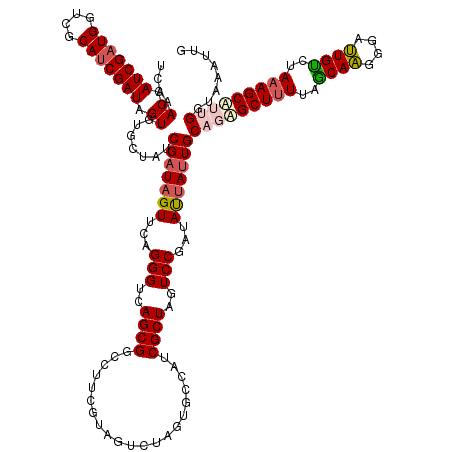
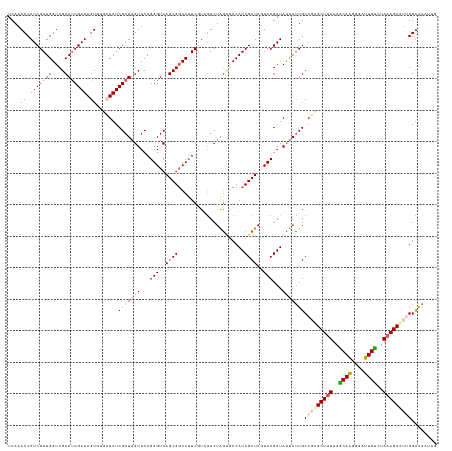
| Location | 16,405,565 – 16,405,701 |
|---|---|
| Length | 136 |
| Sequences | 5 |
| Columns | 136 |
| Reading direction | reverse |
| Mean pairwise identity | 85.29 |
| Shannon entropy | 0.26264 |
| G+C content | 0.46369 |
| Mean single sequence MFE | -38.30 |
| Consensus MFE | -28.18 |
| Energy contribution | -28.70 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.962285 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 16405565 136 - 27905053 GAAUUCACCAAGGCUUUAGACAAUCCCUUGCUAAAAGCCCUGCAAUAAUAUCGGAUUAGCGAUGGCAUUAGACAACGAAGGCCGCUGACCCUGAACUAUCGAUAGCUCACUAUCGAUGCGACCAUCGAUGGUUUGA ........((.((((((((.(((....))))).)))))).))........((((((((.(((((((.((((....(....)...))))........(((((((((....))))))))).).)))))).)))))))) ( -35.70, z-score = -1.88, R) >droSim1.chr3R 22454056 136 + 27517382 CAAUUUACCUACGCUUUAGACAAUCCCUUGCUAAAAGCGCUGCAAUAAUAUCGGACUAGCGAUGGCACUAGACUACGAAGGCCGCUGACCCUGAACUAUCGAUAGCACACUAUCGAUGCGACCAUCGAUGGUUUGA ...........((((((((.(((....))))).))))))...........((((((((.(((((((............(((........)))....(((((((((....))))))))).).)))))).)))))))) ( -35.80, z-score = -2.71, R) >droSec1.super_27 468480 136 + 799419 CAAUUUACCUACGCUUUAUACAAUCCCUUGCUAAAAGCGCUGCAAUAAUAUCGGACUAGCGAUGGCACUAGACUACGAAGGCCGCUGACCCUGAACUAUCGAUAGCACACUAUCGAUGCGACCAUCGAUGGUUUGA ...........((((((...(((....)))...))))))...........((((((((.(((((((............(((........)))....(((((((((....))))))))).).)))))).)))))))) ( -33.90, z-score = -2.34, R) >droYak2.chr3R 5191648 133 - 28832112 ---AUUGCCAAUGCUUUAAACACUUCUGUGUUAAAAGCUGUGCAGUAGUAUCGGAAUAGCGAUGGGUCUAGACAAUGAAGGACGCUGACCCUGAACUAUCGAUACAAUACUAUCGGUGCGACCAUCGAUGGUUUGA ---(((((((..(((((.(((((....))))).))))))).))))).((((((..((((....(((((.((.(......)....)))))))....))))))))))...((((((((((....)))))))))).... ( -44.00, z-score = -3.57, R) >droEre2.scaffold_4770 12479140 136 + 17746568 CCAAUUACCAAUGCUUUAGGCAAUUCCUUGCUGAGAGCUGUGCAAUGGUAUCGGAAUAGCGAUGGGGCCAGAUCAUGACGGCCGCUGACCCUGAACUAUCGAUACAAUACUAUCGAUGCGACCAUCGAUGGUAUGA ............(((((.(((((....))))).)))))(((((.(((((.((((..((((.....((((..........))))))))...))))))))).).))))((((((((((((....)))))))))))).. ( -42.10, z-score = -1.75, R) >consensus CAAUUUACCAACGCUUUAGACAAUCCCUUGCUAAAAGCGCUGCAAUAAUAUCGGAAUAGCGAUGGCACUAGACAACGAAGGCCGCUGACCCUGAACUAUCGAUAGCACACUAUCGAUGCGACCAUCGAUGGUUUGA ...........((((((((.(((....)))))))).)))...........((((((((.((((((.............(((........)))....(((((((((....)))))))))...)))))).)))))))) (-28.18 = -28.70 + 0.52)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:30:20 2011