
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 8,105,931 – 8,106,059 |
| Length | 128 |
| Max. P | 0.500000 |

| Location | 8,105,931 – 8,106,059 |
|---|---|
| Length | 128 |
| Sequences | 12 |
| Columns | 156 |
| Reading direction | forward |
| Mean pairwise identity | 71.02 |
| Shannon entropy | 0.59902 |
| G+C content | 0.43007 |
| Mean single sequence MFE | -30.00 |
| Consensus MFE | -14.80 |
| Energy contribution | -14.79 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.20 |
| Mean z-score | -0.83 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr2L 8105931 128 + 23011544 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGCCUA-------ACUGACUGCCU----GACUGACG--UAACCUGGCAACCGAAACGUGAACG-------------- (((.(((((((((((((...((....))..........(((.((((((((((....)))))).....))))-))))))))).)))-------))))..((((.----.((....)--).....))))........)))....-------------- ( -33.60, z-score = -2.14, R) >droSim1.chr2L_random 424738 128 + 909653 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGCCUA-------ACUGACUGCCU----GACUGACG--UAAGCUGGCAAACGAAACAUGAACG-------------- ....(((((((((((((...((....))..........(((.((((((((((....)))))).....))))-))))))))).)))-------))))..((((.----(.((....--..))).))))...............-------------- ( -32.70, z-score = -1.67, R) >droSec1.super_3 3602340 128 + 7220098 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGCCUA-------ACUGACUGCCU----GACUGACG--UAACCUGGCAAACGAAACAUGAACG-------------- ....(((((((((((((...((....))..........(((.((((((((((....)))))).....))))-))))))))).)))-------))))..((((.----.((....)--).....))))...............-------------- ( -30.80, z-score = -1.63, R) >droYak2.chr2L 10736615 128 + 22324452 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGCCUA-------ACUGACUGCCU----GACUGACG--UAACCUGGUAACUGAAACAUGAACG-------------- ....(((((((((((((..((.(((.(((.........(((.((((((((((....)))))).....))))-)))))).))))).-------..)))))).))----)))))...--.........................-------------- ( -28.80, z-score = -0.95, R) >droEre2.scaffold_4929 17011769 128 + 26641161 CACAUUGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGCCUA-------ACUGACUGCUU----GACUGACG--UAACCUGGUAACCGAAACGUGAACG-------------- (((...(((((((((((..((.(((.(((.........(((.((((((((((....)))))).....))))-)))))).))))).-------..))))))).)----))).....--.....(((...)))....)))....-------------- ( -28.50, z-score = -0.65, R) >droAna3.scaffold_12916 11828873 134 - 16180835 GGCAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCCCAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCCGACUGCCUA-------ACUGACUGCUU----GACUGACA--UAACCUGGUAACAAGACUAAGGACUACCUCA-------- ((..(((((((((((((...((....)).........(((.(((((((((((....)))))).....))))-).)))........-------..))))))).)----)))))...--...((((((......))).)))....))...-------- ( -33.30, z-score = -1.16, R) >dp4.chr4_group4 5626404 120 - 6586962 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGACUA-------ACCGACUCUCCUG--GGCUGACUGCCAGCCUGUUAGCC-------------------------- ......(((((((((((...((....))..........(((.((((((((((....)))))).....))))-)))))))).))))-------))..........(--(((((.....)))))).......-------------------------- ( -32.80, z-score = -1.32, R) >droPer1.super_46 98671 120 + 590583 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGACUA-------ACUGACUCUCCUG--GGCUGACUGCCAGCCUGUUAGCC-------------------------- ....(((((((((((((...((....))..........(((.((((((((((....)))))).....))))-)))))))).))))-------))))........(--(((((.....)))))).......-------------------------- ( -35.50, z-score = -1.96, R) >droWil1.scaffold_180708 6356703 100 + 12563649 CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG-GCCUGACUGACU----------UGACUGACGUA--ACCUGG------------------------------------------- ......(((((((((((...((....))..........(((.((((((((((....)))))).....))))-)))....)))))----------..))))))...--......------------------------------------------- ( -23.20, z-score = -1.14, R) >droGri2.scaffold_15252 13728619 156 + 17193109 CAAGUAGCUGGCAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUGCGCCCCUCCACCCCCCGGUACACCACCUCUCCCUCCUAUUAACCAACCACUUGACUGACGUAACCUGGAGAGAGCAAAGUCAGUG (((((.(.(((.........((....))...((((((.(((.((((((((((....)))))).....)))).)))............(((.....))).........))))))))).).)))))((((((.....((......))....)))))). ( -32.60, z-score = -0.25, R) >droMoj3.scaffold_6500 21242539 120 + 32352404 ----UAGUUGGCAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUGCCCCCCU--------------------------------UAACCAAGCACUUGACUGACGUAACCUGGAGAGCAUAGCAUAGGCG ----((((((((.((((.((((....)).)).))))...)).)))))).........(((((.(((((((..((...(--------------------------------((....(((....).))....)))...))..)))))))..))))). ( -21.90, z-score = 1.66, R) >droVir3.scaffold_12963 16303169 140 + 20206255 ----UAGUUGGCAGUUAUAAGAGCAAUCAUUUUAAUAUGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUGCCGACCGCAAGCCC-------ACCACGUGCCCCGCCCCCUAACCGAACACUUGACUGACGUAACCUGGAGAGCAGAGCAU----- ----..(((((((((((...((....))..........(((((...((((((....))))))......((((.....))))....-------.....)))))....................)))))).).))))(((.....))).....----- ( -26.30, z-score = 1.22, R) >consensus CACAUAGUUAGUAGUUAUAAGAGCAAUCAUUUUAAUACGGCACAGCUAAGCACUUUUGCUUAAUUAUGUUG_GCCUGACUGCCUA_______ACUGACUGCCU____GACUGACG__UAACCUGGCAACCGAAACAUGAACG______________ ......(((((..((..(((((.......)))))..))(((.((((((((((....)))))).....)))).)))..................................))))).......................................... (-14.80 = -14.79 + -0.01)

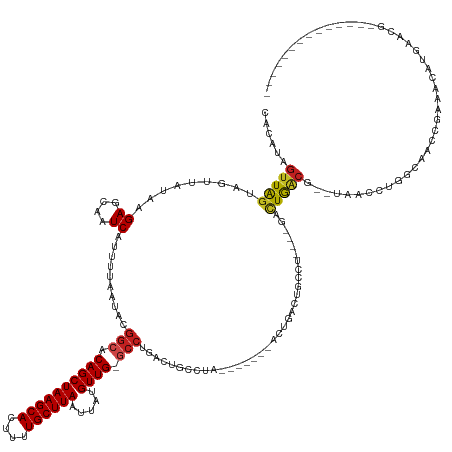

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:25:04 2011