
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 16,041,491 – 16,041,591 |
| Length | 100 |
| Max. P | 0.753280 |

| Location | 16,041,491 – 16,041,591 |
|---|---|
| Length | 100 |
| Sequences | 10 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 76.95 |
| Shannon entropy | 0.46739 |
| G+C content | 0.49480 |
| Mean single sequence MFE | -34.34 |
| Consensus MFE | -11.79 |
| Energy contribution | -13.44 |
| Covariance contribution | 1.65 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.753280 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 16041491 100 + 27905053 ----CGUUUUGUGGUAGCG---AAUGGCCACCGCAAACAAGGAC-UGCA--AACAGUUGCAAUCCGC-UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGCAGCCAAU ----.((((.(((((.((.---....)).)))))))))..((.(-((((--....(((((((.....-)))))))...((((((......)))))).....)))))))... ( -35.00, z-score = -2.31, R) >droSim1.chr3R 22076357 100 - 27517382 ----CGUUUUGCGGUGGCG---AAUGGCCACCGCAAACAAGGAC-UGCA--AACAGUUGCAAUCCGC-UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGCAGCCAAU ----.((((.((((((((.---....))))))))))))..((.(-((((--....(((((((.....-)))))))...((((((......)))))).....)))))))... ( -43.40, z-score = -4.36, R) >droSec1.super_27 108559 100 - 799419 ----CGUUUUGCGGUGGCG---AAUGGCCACCGCAAACAAGGAC-UGCA--AACAGUUGCAAUCCGC-UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGCAGCCAAU ----.((((.((((((((.---....))))))))))))..((.(-((((--....(((((((.....-)))))))...((((((......)))))).....)))))))... ( -43.40, z-score = -4.36, R) >droYak2.chr3R_random 312675 100 + 3277457 ----CGUUUUGCGGUGGCG---AAUGGCCACCGCAAACAAGGAC-UGCA--AACAGUUGCAAUCCGC-UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGCAGCCAAU ----.((((.((((((((.---....))))))))))))..((.(-((((--....(((((((.....-)))))))...((((((......)))))).....)))))))... ( -43.40, z-score = -4.36, R) >droEre2.scaffold_4770 12115015 100 - 17746568 ----CGUUUUGCGGUGGCG---AAUGGCCACCGCAAACAAGGAC-UGCA--AACAGUUGCAAUCCGC-UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGCAGCCAAU ----.((((.((((((((.---....))))))))))))..((.(-((((--....(((((((.....-)))))))...((((((......)))))).....)))))))... ( -43.40, z-score = -4.36, R) >droAna3.scaffold_13340 2591946 101 + 23697760 ----CAGUUUGCGGUGGUGG--AAUGGCCACCGCAAACAAGGAC-AGCA--AACAGUUGCAAUCCGC-UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGGAGCCAAU ----..((((((((((((..--....))))))))))))..(((.-.(((--(....))))..)))..-(((...((((((((((......)))))).....))))..))). ( -40.80, z-score = -3.49, R) >dp4.chr2 3145912 109 - 30794189 CCAACGUUUUGUGGCUGUGCGGAUGGGCUACCGCAAACAAGGACAUGCA--AACAGUAGGAAUCCCCCUUGUAAUCCAGCAAGGAUUUGACUUUGCACACAUGCAGGCAGU ........((((.((((((.((((((....))....((((((...(((.--....)))((....)))))))).)))).((((((......)))))).)))).))..)))). ( -29.60, z-score = 0.58, R) >droPer1.super_6 3169145 109 - 6141320 CCAACGUUUUGUGGCUGUGCGGAUGGGCUACCGCAAACAAGGACAUGCA--AACAGUAGGAAUCCCCCUUGUAAUCCAGCAAGGAUUUGACUUUGCACACAUGCAGGCAGU ........((((.((((((.((((((....))....((((((...(((.--....)))((....)))))))).)))).((((((......)))))).)))).))..)))). ( -29.60, z-score = 0.58, R) >droGri2.scaffold_15074 502252 85 - 7742996 ---------------------UAAUAACCACAGCAAACAACAC----CAGCAGCAGCAGAAAUCCGC-CUGUAAUCCUGAAAGGAUUUAACUUGGCAUACAUGCAACUCAU ---------------------...........(((........----..((....))........((-(.(((((((.....)))))..))..))).....)))....... ( -14.30, z-score = -0.94, R) >droMoj3.scaffold_6540 14229858 88 - 34148556 ---------------------UAAUAGCCACAGAAAACAAAACACAACAGCAGCAGCAAAAAUCCGG-CUGUAAUCCUGCCAGGAUUUGACUUGCUGUACAAGCAGAAUU- ---------------------............................(((((((...((((((((-(.........))).))))))...)))))))............- ( -20.50, z-score = -1.98, R) >consensus ____CGUUUUGCGGUGGCG___AAUGGCCACCGCAAACAAGGAC_UGCA__AACAGUUGCAAUCCGC_UUGUAAUCCAGCCAGGAUUUGACUUGGCACACAUGCAGCCAAU .......(((((((..((........))..)))))))...............................(((((.....((((((......)))))).....)))))..... (-11.79 = -13.44 + 1.65)


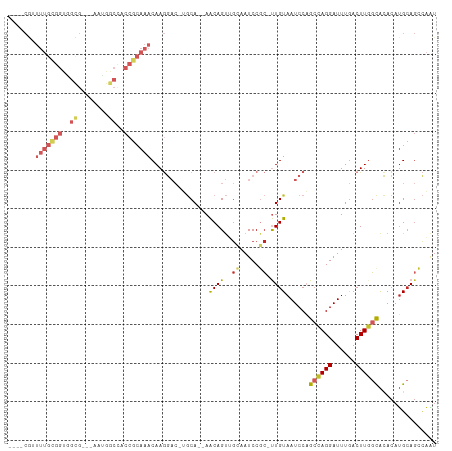
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:29:18 2011