
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 15,762,493 – 15,762,591 |
| Length | 98 |
| Max. P | 0.749457 |

| Location | 15,762,493 – 15,762,591 |
|---|---|
| Length | 98 |
| Sequences | 10 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 64.06 |
| Shannon entropy | 0.72650 |
| G+C content | 0.44344 |
| Mean single sequence MFE | -22.25 |
| Consensus MFE | -8.16 |
| Energy contribution | -8.48 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.749457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 15762493 98 + 27905053 -UUUCCCAUGCCUGUCCAUGCAUAACUAC---CAUUCAUUG---AACGC-GCUAUAAGGUAACGCGAUC----UAAUGAUGUUUCCAUGGGAUCCAUUCUCACUUCUCCU--- -..(((((((..(((....))).......---...((((((---(.(((-(.(((...))).)))).))----.)))))......)))))))..................--- ( -19.90, z-score = -1.57, R) >droSim1.chr3R 21805568 98 - 27517382 -UUUCCCAUGCCUGUCCAUGCAUAACUGC---CAUUCAUUG---AACGC-GCUAUAAGGUAACGCGAUC----UAAUGAUGUUUCCAUGGGAUCCAUUCUUCCUAUUCCU--- -..(((((((.........(((....)))---...((((((---(.(((-(.(((...))).)))).))----.)))))......)))))))..................--- ( -21.70, z-score = -1.72, R) >droSec1.super_25 812481 98 - 827797 -UUUCCCAUGCCUGUCCAUGCAUAACUGC---CAUUCAUUG---AACGC-GCUAUAAGGUAACGCGAUC----UAAUGAUGUUUCCAUGGGAUCCAUUCUUCCUAUUCCU--- -..(((((((.........(((....)))---...((((((---(.(((-(.(((...))).)))).))----.)))))......)))))))..................--- ( -21.70, z-score = -1.72, R) >droYak2.chr3R 4536992 95 + 28832112 -UUUCCCAUGCCUGGCCAUGCAUAAAUGC---UAUUCAUUG---AACGC-GCUAUAAGGUAACGCGAUC-------UGAUGGUUCCAUGGGAUCCAUUCUCCCUUCUUCG--- -..(((((((...(((((((((....)))---........(---(.(((-(.(((...))).)))).))-------..)))))).)))))))..................--- ( -24.20, z-score = -1.35, R) >droEre2.scaffold_4770 11848101 80 - 17746568 -UUUCCCAUGCCUGUCCAUGCAUAACUGC---UAUUCAUUG---AACGC-GCUGUAAGGUGAGGCGGUC----UAAUGAUGGUUCCAUGGGA--------------------- -..(((((((.....((((.(((.(((((---(..(((((.---.((..-...))..))))))))))).----..)))))))...)))))))--------------------- ( -25.40, z-score = -1.32, R) >droAna3.scaffold_13340 2343426 102 + 23697760 -UUUCCCAUGCCUGCUCAUGCAUAACCGA---UGCUUAAUG---AAUGC-GCUAAAAAGUAACGCGAUC-UAAUAAUGAUGGCACCUACGGAUCCAUUUUCUGUUAAUUCA-- -.......((((...((((((((.....)---))).....(---(.(((-(.((.....)).)))).))-.....)))).))))...(((((.......))))).......-- ( -17.80, z-score = -0.23, R) >dp4.chr2 2862264 106 - 30794189 -UUUCCCAGGCCUGCUCGUGCAUAACCGA---UGUUCAUGG--CAAUGCCGCUAAAAAGUAAUGCGAUC-UAAUGAUGGCACCACCAUGGGACCCAUUCAAUUUCCCUAACCU -..((((((((.(((.((((.(((.....---))).)))))--))..)))........((..(((.(((-....))).)))..))..)))))..................... ( -25.90, z-score = -0.94, R) >droPer1.super_6 2881531 106 - 6141320 -UUUCCCAGGCCUGCUCGUGCAUAACCGA---UGUUCAUGG--CAAUGCCGCUAAAAAGUAAUGCGAUC-UAAUGAUGGCACCACCAUGGGAUCCAUUCAAUUUCCCUAACCU -..((((((((.(((.((((.(((.....---))).)))))--))..)))........((..(((.(((-....))).)))..))..)))))..................... ( -26.00, z-score = -0.91, R) >droWil1.scaffold_181108 660450 88 - 4707319 -UUUCCCAUGCCUGGUCAUGCAUAAACUG---ACUUUAACAU-UUAUUCUAUUGUAGGG-AUCAUAUUC------AUGGGAUCCCAUUCACAUGUCUUCU------------- -..(((((((..(((((.((((.......---..........-.........))))..)-))))....)------))))))...................------------- ( -16.27, z-score = -0.26, R) >droGri2.scaffold_15074 2352128 96 - 7742996 AUUUCCCAUGCCUGAUCAGGCAUAACCAAAUUUAAUUAGCAUUCAAUGC-GCCAAAA-GUAGCGCAAAGACAGAAGAGAUGGCAACACGGGAUCCAUU--------------- ...((((((((((....))))))...............((......(((-((.....-...)))))....((.......)))).....))))......--------------- ( -23.60, z-score = -2.14, R) >consensus _UUUCCCAUGCCUGUCCAUGCAUAACCGC___UAUUCAUUG___AACGC_GCUAUAAGGUAACGCGAUC____UAAUGAUGGCUCCAUGGGAUCCAUUCU__CU_CUUC____ ...((((((((........)))).......................(((.(((....)))...)))......................))))..................... ( -8.16 = -8.48 + 0.32)


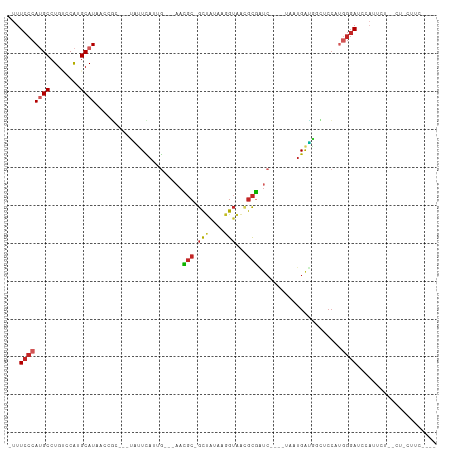
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:28:44 2011