
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 15,337,785 – 15,337,888 |
| Length | 103 |
| Max. P | 0.529432 |

| Location | 15,337,785 – 15,337,888 |
|---|---|
| Length | 103 |
| Sequences | 8 |
| Columns | 121 |
| Reading direction | reverse |
| Mean pairwise identity | 66.63 |
| Shannon entropy | 0.60945 |
| G+C content | 0.49660 |
| Mean single sequence MFE | -27.93 |
| Consensus MFE | -13.59 |
| Energy contribution | -12.76 |
| Covariance contribution | -0.83 |
| Combinations/Pair | 1.59 |
| Mean z-score | -0.81 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.529432 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
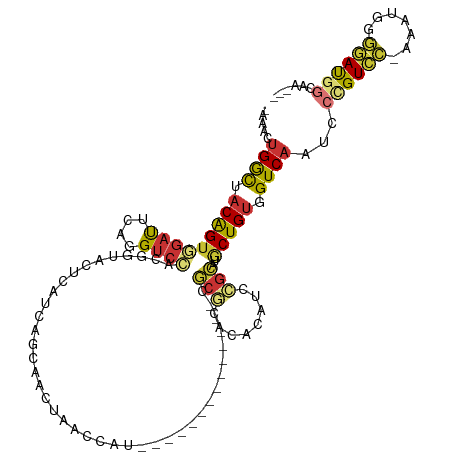
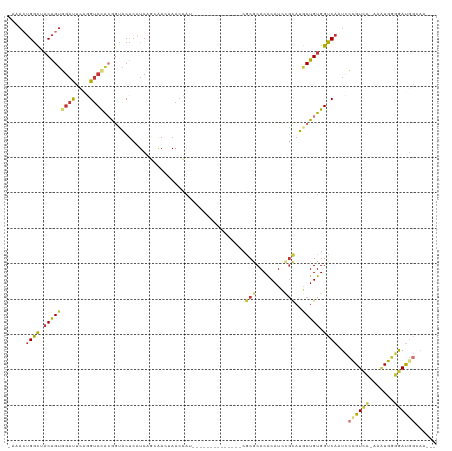
>dm3.chr3R 15337785 103 - 27905053 AAAACUGGCUACAGUGGAUUCAGGUCCGCGGUACUAAUCACCAACUAACCAU--------------CGCGCACACAUCCGCAUGCUGUGGUCAAUCCCGUCC-AAAUGGGGAUGGCAA--- .....((((((((((((((....((.((((((..................))--------------)))).))..))))....))))))))))...((((((-......))))))...--- ( -32.47, z-score = -1.38, R) >droSim1.chr3R 21379782 102 + 27517382 -AAACUGGCUACAGUGGAUUCAGGUCCACGGUACUCAUCAGCAACUAACCAU--------------CGCGCACACAUCCGCAUGCUGUGGUCAAUCCCGUCC-AAAUGGGGAUGGAAA--- -....((((((((((((((....))))..(((...............)))..--------------.(((........)))..))))))))))...((((((-......))))))...--- ( -30.86, z-score = -1.13, R) >droSec1.super_25 389278 102 + 827797 -AAACUGGCUACAGUGGAUUCAGGUCCACGGUACUCAUCAGCAACUAACCAU--------------CGCGCACGCAUCCGCAUGCUGUGGUCAAUCCCGUCC-AAAUGGGGAUGGAAA--- -....((((((((((((((....))))))(((...............)))..--------------...(((.((....)).)))))))))))...((((((-......))))))...--- ( -33.06, z-score = -1.50, R) >droYak2.chr3R 4077282 99 - 28832112 -AAAAUGGCUACAGUGGAUUUAGGUCCACGGUACUCAUCAG-GACUAACCAU--------------CGCGCACACAUCCGCAUGCUGUGGUCAAUCCUGUCC---GUGGGAAUGGCAA--- -......((((..((((((....))))))....((((((((-((.(.(((((--------------.((((........))..)).))))).).)))))...---)))))..))))..--- ( -31.50, z-score = -1.12, R) >droEre2.scaffold_4770 11442533 102 + 17746568 -AAACUGGCUACAGUGGAUUCAGGUCCACGGUACUCAUCGGGAACUAACCAU--------------CGCGCACACAUCCGCAUGCUGUGGUCAAUCCCGUCC-AAAUGGGGAUGGCAG--- -....((((((((((((.((.((.(((.((((....))))))).))))))..--------------.(((........)))..))))))))))...((((((-......))))))...--- ( -33.70, z-score = -1.30, R) >droAna3.scaffold_13340 8635284 94 + 23697760 ------GAC--CGGCAACUCUGCGACUACAGUGCCCAUCA--AA--AAGCAG--------------GACACACACAUCUACACACUGUGGUCAUUCCUGUCC-AGCUGCCGAUGGCGGAAA ------..(--(((((...(((......))))))(((((.--..--.(((.(--------------((((.......((((.....)))).......)))))-.)))...))))))))... ( -27.04, z-score = -1.15, R) >dp4.chr2 29817624 111 - 30794189 -AAACUGGUAACAGUCGACAACAGUCCA-AACGCUCAUCAAAAAAAAACUUCACACAGAUACAGCACACACACACAUUCGUACACUGUGGUCAUUUGUGUUUGAAAUGCAAAC-------- -..((((....)))).(((....)))..-...((...((((...........((((((((...((.((((...((....))....)))))).))))))))))))...))....-------- ( -17.40, z-score = 0.56, R) >droPer1.super_6 5186961 111 - 6141320 -AAACUGGUAACAGUCGACAACAGUCCA-AACGCUCAUCAAAAAAAAACUUCACACAGAUACAGCACACACACACAUUCGUACACUGUGGUCAUUUGUGUUUGAAAUGCAAAC-------- -..((((....)))).(((....)))..-...((...((((...........((((((((...((.((((...((....))....)))))).))))))))))))...))....-------- ( -17.40, z-score = 0.56, R) >consensus _AAACUGGCUACAGUGGAUUCAGGUCCACGGUACUCAUCAGCAACUAACCAU______________CGCGCACACAUCCGCAUGCUGUGGUCAAUCCCGUCC_AAAUGGGGAUGGCAA___ .....((((.(((((((((....))))........................................(((........)))..))))).)))).((((((.....)))))).......... (-13.59 = -12.76 + -0.83)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:27:50 2011