| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,962,603 – 7,962,700 |
| Length | 97 |
| Max. P | 0.993804 |

| Location | 7,962,603 – 7,962,700 |
|---|---|
| Length | 97 |
| Sequences | 8 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 71.85 |
| Shannon entropy | 0.54788 |
| G+C content | 0.46994 |
| Mean single sequence MFE | -22.99 |
| Consensus MFE | -11.13 |
| Energy contribution | -11.30 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.64 |
| SVM RNA-class probability | 0.993804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
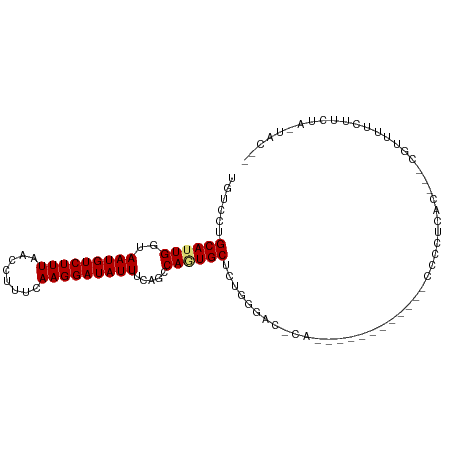
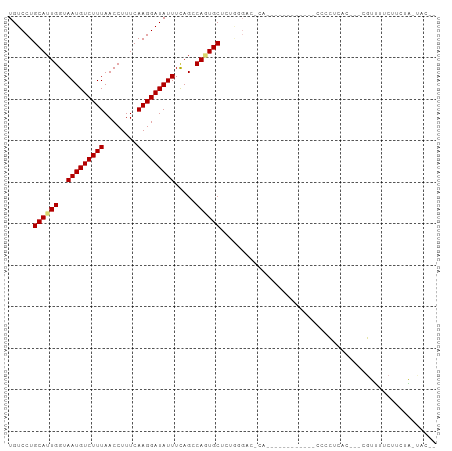
>dm3.chr2L 7962603 97 + 23011544 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUUAGCCAGUGCUCUGGGACGCACCACUUGCUACCCCCCUAAC---CGUUUUCUUCUA-UAC-- .((((((((((((((((((((((........)))))))))...))))))))...)))))(((.....)))............---............-...-- ( -25.90, z-score = -3.28, R) >droAna3.scaffold_12943 3691293 79 - 5039921 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGGCAAUGCUCGGAGAG----------------CGCCCUC---CGCCUUCCUGUG-C---- ......((((((.((((((((((........)))))))))...).))))))..((((.(----------------((.....---))).))))....-.---- ( -20.30, z-score = -0.63, R) >droEre2.scaffold_4929 16883620 86 + 26641161 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGCCAGUGCUCUGGGACGCAC-----------CCCCUUAG---CGUUUUCUGCUA-UAC-- .((((((((((((((((((((((........)))))))))...))))))))...)))))....-----------.....(((---((.....)))))-...-- ( -27.50, z-score = -3.49, R) >droYak2.chr2L 17392735 97 - 22324452 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGCCAGUGCUCUGGGGGACACGCACCACCUACCCCCUCGC---CGUUUUCUUCUA-UAC-- ......(((((((((((((((((........)))))))))...))))))))...(((((..............)))))....---............-...-- ( -26.14, z-score = -2.06, R) >droSec1.super_3 3474693 73 + 7220098 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGCCAGUGC------------------------CCCUAAC---CGUUUUCUUCUA-UAC-- ......(((((((((((((((((........)))))))))...))))))))------------------------.......---............-...-- ( -17.10, z-score = -3.37, R) >droSim1.chr2L 7756588 74 + 22036055 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGCCAGUGC-----------------------CCCCUAAC---CGUUUUCUUCUA-UAC-- ......(((((((((((((((((........)))))))))...))))))))-----------------------........---............-...-- ( -17.10, z-score = -3.49, R) >dp4.chr4_group3 6387282 103 - 11692001 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGGCAUUGCCAUAUGCCCCAUGAACCAUGCGCCCCUCCACAGCCACUAUCCUCUGGCACCC ......(((((((((((((((.((.(((....))).))....))))))))))).))))..((((...))))............((((........)))).... ( -25.50, z-score = -1.73, R) >droPer1.super_1 3487370 103 - 10282868 UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGGCAUUGCCUCAUGCCCCAUAACCCAUGCGCCCCUUCACAGCCACUAUCCUCUGGCACCC ......((((.((((((((((((........)))))))))...(((((......))))).....))).))))...........((((........)))).... ( -24.40, z-score = -1.91, R) >consensus UGUCCUGCAUUGGUAAUGUCUUUAACCUUUCAAGGAUAUUUCAGCCAGUGCUCUGGGAC_CA____________CCCCUCAC___CGUUUUCUUCUA_UAC__ ......((((((..(((((((((........))))))))).....)))))).................................................... (-11.13 = -11.30 + 0.17)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:24:52 2011