
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,957,956 – 7,958,065 |
| Length | 109 |
| Max. P | 0.984845 |

| Location | 7,957,956 – 7,958,065 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 121 |
| Reading direction | forward |
| Mean pairwise identity | 66.67 |
| Shannon entropy | 0.51933 |
| G+C content | 0.28077 |
| Mean single sequence MFE | -23.09 |
| Consensus MFE | -11.24 |
| Energy contribution | -16.18 |
| Covariance contribution | 4.94 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.18 |
| SVM RNA-class probability | 0.984845 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
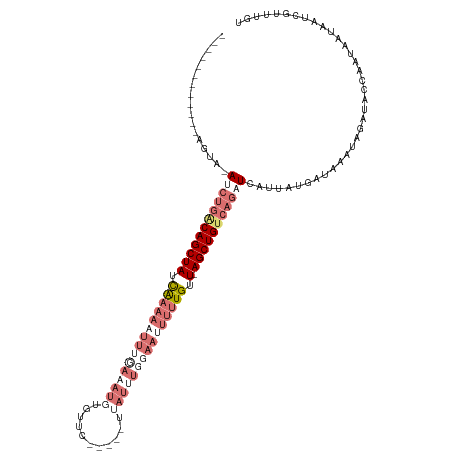
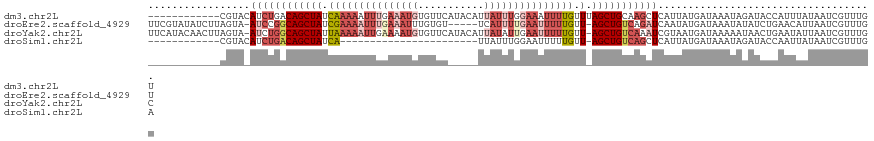
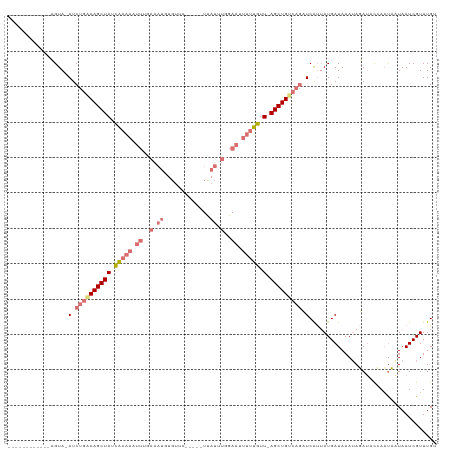
>dm3.chr2L 7957956 109 + 23011544 ------------CGUACAUCUGACAGCUAUCAAAAAUUUGAAAUGUGUUCAUACAUUAUUUGGAAAUUUUGUUUAGCUGCAAGCUCAUUAUGAUAAAUAGAUACCAUUUAUAAUCGUUUGU ------------...........((((((.(((((.(((.((((((((....))).))))).))).)))))..))))))(((((..(((((((..............))))))).))))). ( -21.74, z-score = -1.44, R) >droEre2.scaffold_4929 16880385 114 + 26641161 UUCGUAUAUCUUAGUA-AUCCGGCAGCUAUCGAAAAUUUGAAAUUUGUGU-----UCAUUUUGAAUUUUUGUU-AGCUGUCAGAUCAAUAUGAUAAAUAUAUCUGAACAUUAAUCGUUUGU ......((((.((...-(((.((((((((.((((((((..((((......-----..))))..)))))))).)-))))))).)))...)).)))).........((((.......)))).. ( -28.50, z-score = -3.62, R) >droYak2.chr2L 17389301 119 - 22324452 UUCAUACAACUUAGUA-AUCUGGCAGCUAUUAAAAAUUGAAAAUGUGUUCAUACAUUAUAUUGAAUUUUUGUU-AGCUGUCAAAUCGUAAUGAUAAAAAUAACUGAAUAUUAAUCGUUUGC ..((.((...((((((-.(((((((((((.((((((((.((((((((....))))))...)).)))))))).)-)))))))).(((.....)))..........)).))))))..)).)). ( -24.10, z-score = -2.37, R) >droSim1.chr2L 7753379 85 + 22036055 ------------CGUACAUCUGACAGCUAUCA-----------------------UUAUUUGGAAUUUUUGUU-AGCUGUCAGCUCAUUAUGAUAAAUAGAUACCAAUUAUAAUCGUUUGA ------------((.((..((((((((((.((-----------------------..(........)..)).)-)))))))))...((((((((............)))))))).)).)). ( -18.00, z-score = -1.90, R) >consensus ____________AGUA_AUCUGACAGCUAUCAAAAAUUUGAAAUGUGUUC_____UUAUUUGGAAUUUUUGUU_AGCUGUCAGAUCAUUAUGAUAAAUAGAUACCAAUAAUAAUCGUUUGU ...................(((((((((..(((((((((((((((...........)))))))))))))))...)))))))))...................................... (-11.24 = -16.18 + 4.94)

| Location | 7,957,956 – 7,958,065 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 121 |
| Reading direction | reverse |
| Mean pairwise identity | 66.67 |
| Shannon entropy | 0.51933 |
| G+C content | 0.28077 |
| Mean single sequence MFE | -20.01 |
| Consensus MFE | -7.72 |
| Energy contribution | -10.79 |
| Covariance contribution | 3.06 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.913522 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
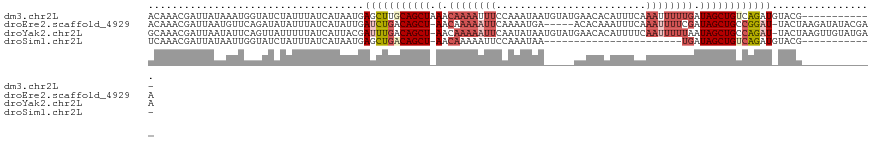
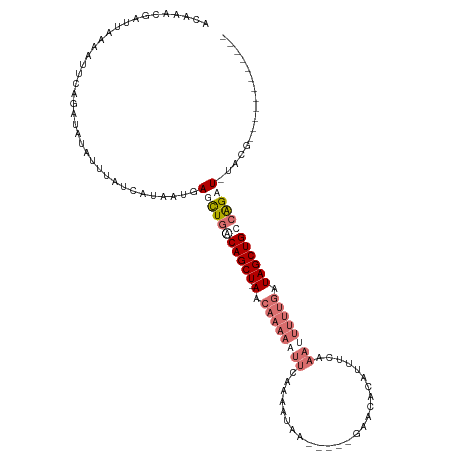
>dm3.chr2L 7957956 109 - 23011544 ACAAACGAUUAUAAAUGGUAUCUAUUUAUCAUAAUGAGCUUGCAGCUAAACAAAAUUUCCAAAUAAUGUAUGAACACAUUUCAAAUUUUUGAUAGCUGUCAGAUGUACG------------ (((...........((((((......))))))......((.(((((((...(((((((......(((((......)))))..)))))))...))))))).)).)))...------------ ( -17.70, z-score = -0.82, R) >droEre2.scaffold_4929 16880385 114 - 26641161 ACAAACGAUUAAUGUUCAGAUAUAUUUAUCAUAUUGAUCUGACAGCU-AACAAAAAUUCAAAAUGA-----ACACAAAUUUCAAAUUUUCGAUAGCUGCCGGAU-UACUAAGAUAUACGAA .....((...(((((......)))))((((.((.(((((((.(((((-(.(.((((((..((((..-----......))))..)))))).).)))))).)))))-)).)).))))..)).. ( -23.40, z-score = -3.79, R) >droYak2.chr2L 17389301 119 + 22324452 GCAAACGAUUAAUAUUCAGUUAUUUUUAUCAUUACGAUUUGACAGCU-AACAAAAAUUCAAUAUAAUGUAUGAACACAUUUUCAAUUUUUAAUAGCUGCCAGAU-UACUAAGUUGUAUGAA ((((..(((.((((......))))...))).....((((((.(((((-(..(((((((......(((((......)))))...)))))))..)))))).)))))-)......))))..... ( -20.30, z-score = -1.71, R) >droSim1.chr2L 7753379 85 - 22036055 UCAAACGAUUAUAAUUGGUAUCUAUUUAUCAUAAUGAGCUGACAGCU-AACAAAAAUUCCAAAUAA-----------------------UGAUAGCUGUCAGAUGUACG------------ ....(((.((((...(((((......)))))..)))).(((((((((-(.((..............-----------------------)).)))))))))).)))...------------ ( -18.64, z-score = -2.93, R) >consensus ACAAACGAUUAAAAUUCAGAUAUAUUUAUCAUAAUGAGCUGACAGCU_AACAAAAAUUCAAAAUAA_____GAACACAUUUCAAAUUUUUGAUAGCUGCCAGAU_UACG____________ ....................................(((((((((((...((((((((.........................))))))))..)))))))))))................. ( -7.72 = -10.79 + 3.06)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:24:51 2011