
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 15,077,100 – 15,077,202 |
| Length | 102 |
| Max. P | 0.871521 |

| Location | 15,077,100 – 15,077,202 |
|---|---|
| Length | 102 |
| Sequences | 10 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 96.50 |
| Shannon entropy | 0.07806 |
| G+C content | 0.39275 |
| Mean single sequence MFE | -21.48 |
| Consensus MFE | -20.46 |
| Energy contribution | -20.56 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.871521 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 15077100 102 - 27905053 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.56, R) >droSim1.chr3R 21112841 102 + 27517382 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.56, R) >droSec1.super_25 138205 102 + 827797 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.56, R) >droYak2.chr3R 3780982 102 - 28832112 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUUAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.53, R) >droEre2.scaffold_4770 11191731 107 + 17746568 -GCAAGUCGCAAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA -((...((((..(((....)))..)))).(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -24.70, z-score = -2.45, R) >droAna3.scaffold_13340 8388114 102 + 23697760 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCGGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.29, R) >dp4.chr2 29525117 102 - 30794189 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.56, R) >droPer1.super_6 4895685 102 - 6141320 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.56, R) >droWil1.scaffold_181130 12839525 102 + 16660200 ------GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAAA ------((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.10, z-score = -1.72, R) >droVir3.scaffold_13047 2654914 108 + 19223366 CGACAAGCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGU ......((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... ( -21.30, z-score = -1.42, R) >consensus ______GCAACAGGCGCAAGUCAAGUGAAUGGCCAAUGACCACCAAAUUGGCAGUAAAGCAUACAAAAAUAACUUCAAAUGGUCAAAAGGUUAACAAAAUCAAGCAGA ......((....(((....))).......(((((..((((((((.....))..(((.....)))...............))))))...)))))..........))... (-20.46 = -20.56 + 0.10)

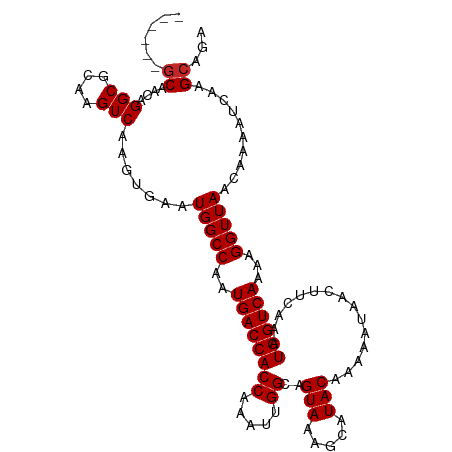
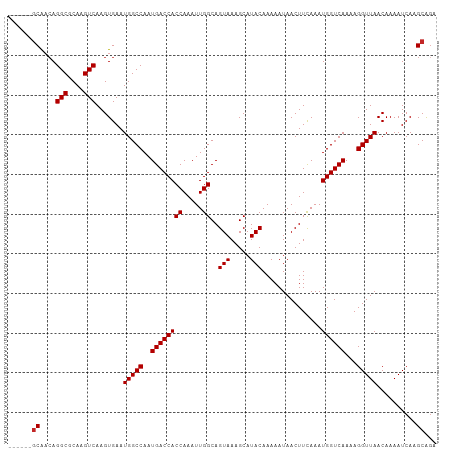
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:27:14 2011