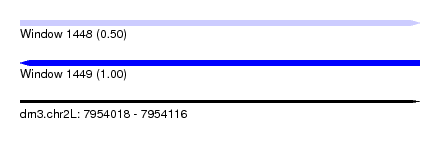

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,954,018 – 7,954,116 |
| Length | 98 |
| Max. P | 0.996656 |

| Location | 7,954,018 – 7,954,116 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 71.23 |
| Shannon entropy | 0.48366 |
| G+C content | 0.58302 |
| Mean single sequence MFE | -18.26 |
| Consensus MFE | -9.79 |
| Energy contribution | -10.19 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
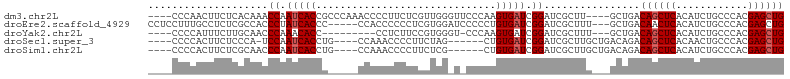
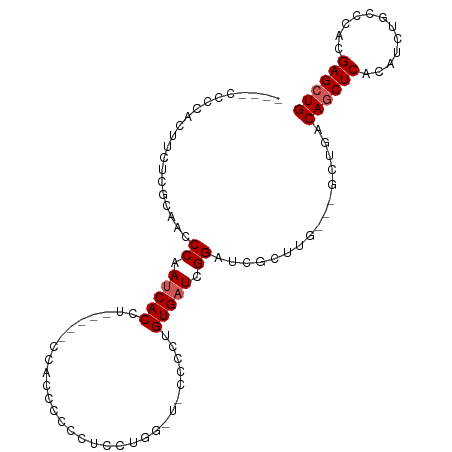
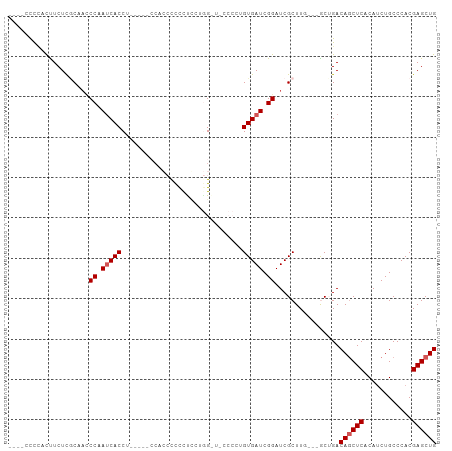
>dm3.chr2L 7954018 98 + 23011544 ----CCCAACUUCUCACAAACCAAUCACCGCCCAAACCCCUUCUCGUUGGGUUCCCAAGUGAUCGGAUCGCUU----GCUGACAGCUCACAUCUGCCCACGAGCUG ----....................(((..((((((...........))))))...((((((((...)))))))----).)))((((((............)))))) ( -20.40, z-score = -1.16, R) >droEre2.scaffold_4929 16876438 98 + 26641161 CCUCCUUUGCCUCUCGCCACCCUAUCACCC-----CCACCCCCCUCGUGGAUCCCCCUGUGAUCGGAUCGCUUU---GCUGACAACUCACAUCUGCCCACGAGCUG ..............................-----........(((((((((((..........)))))((..(---(.(((....)))))...)).))))))... ( -16.70, z-score = -1.40, R) >droYak2.chr2L 17385201 89 - 22324452 ----CCCCAUUUCUUGCAACCCAAACACC---------CCUCUUCCGUGGGU-CCCAAGUGAUCGGAUCGCUUU---GCUGACAGCUCACAUCUGCCCACGAGCUG ----.((.(((.((((..(((((......---------.........)))))-..)))).))).))........---.....((((((............)))))) ( -20.26, z-score = -1.07, R) >droSec1.super_3 3467690 91 + 7220098 ----CCCCACUUCUCCCA-UCCAAUCACCUG----CCAAACCCCUUCUAG------CUGUGAUCGGAUCGCUUGCUGACAGACAGCUCACAACUGCCCACGAGCUG ----.............(-(((.(((((..(----(.............)------).))))).))))..............((((((............)))))) ( -17.12, z-score = -1.39, R) >droSim1.chr2L 7749520 92 + 22036055 ----CCCCACUUCUCGCAACCCAAUCACCUG----CCAAACCCCUUCUCG------CUGUGAUCGGAUCGCUUGCUGACAGACAGCUCACAUCUGCCCACGAGCUG ----...........((((.((.(((((..(----(.............)------).))))).)).....)))).......((((((............)))))) ( -16.82, z-score = -0.89, R) >consensus ____CCCCACUUCUCGCAACCCAAUCACCU_____CCACCCCCCUCCUGG_U_CCCCUGUGAUCGGAUCGCUUG___GCUGACAGCUCACAUCUGCCCACGAGCUG ....................((.(((((..............................))))).))................((((((............)))))) ( -9.79 = -10.19 + 0.40)
| Location | 7,954,018 – 7,954,116 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 71.23 |
| Shannon entropy | 0.48366 |
| G+C content | 0.58302 |
| Mean single sequence MFE | -33.58 |
| Consensus MFE | -21.67 |
| Energy contribution | -22.47 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.96 |
| SVM RNA-class probability | 0.996656 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
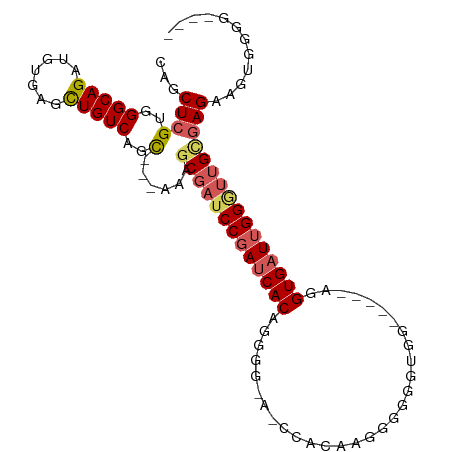
>dm3.chr2L 7954018 98 - 23011544 CAGCUCGUGGGCAGAUGUGAGCUGUCAGC----AAGCGAUCCGAUCACUUGGGAACCCAACGAGAAGGGGUUUGGGCGGUGAUUGGUUUGUGAGAAGUUGGG---- ((((((..(......)..))))))(((((----..((((.(((((((((...((((((.........))))))....))))))))).)))).....))))).---- ( -34.80, z-score = -1.90, R) >droEre2.scaffold_4929 16876438 98 - 26641161 CAGCUCGUGGGCAGAUGUGAGUUGUCAGC---AAAGCGAUCCGAUCACAGGGGGAUCCACGAGGGGGGUGG-----GGGUGAUAGGGUGGCGAGAGGCAAAGGAGG ((((((..(......)..))))))...((---...((.((((.(((((.......(((((.......))))-----).))))).)))).)).....))........ ( -29.00, z-score = -1.79, R) >droYak2.chr2L 17385201 89 + 22324452 CAGCUCGUGGGCAGAUGUGAGCUGUCAGC---AAAGCGAUCCGAUCACUUGGG-ACCCACGGAAGAGG---------GGUGUUUGGGUUGCAAGAAAUGGGG---- ...(((((.(((((.......))))).))---...(((((((((.(((.....-.(((.(....).))---------)))).))))))))).......))).---- ( -30.80, z-score = -1.47, R) >droSec1.super_3 3467690 91 - 7220098 CAGCUCGUGGGCAGUUGUGAGCUGUCUGUCAGCAAGCGAUCCGAUCACAG------CUAGAAGGGGUUUGG----CAGGUGAUUGGA-UGGGAGAAGUGGGG---- ..(((.(..((((((.....))))))..).)))...(.((((((((((.(------(((((.....)))))----)..)))))))))-).)...........---- ( -35.80, z-score = -3.45, R) >droSim1.chr2L 7749520 92 - 22036055 CAGCUCGUGGGCAGAUGUGAGCUGUCUGUCAGCAAGCGAUCCGAUCACAG------CGAGAAGGGGUUUGG----CAGGUGAUUGGGUUGCGAGAAGUGGGG---- ...(((((.(((((((.......))))))).))..(((((((((((((.(------(.(((.....))).)----)..))))))))))))))))........---- ( -37.50, z-score = -3.63, R) >consensus CAGCUCGUGGGCAGAUGUGAGCUGUCAGC___AAAGCGAUCCGAUCACAGGGG_A_CCACAAGGGGGGUGG_____AGGUGAUUGGGUUGCGAGAAGUGGGG____ ((((((..(......)..))))))...........(((((((((((((..............................)))))))))))))............... (-21.67 = -22.47 + 0.80)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:24:49 2011