
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,941,151 – 7,941,245 |
| Length | 94 |
| Max. P | 0.727522 |

| Location | 7,941,151 – 7,941,245 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 73.52 |
| Shannon entropy | 0.49509 |
| G+C content | 0.56983 |
| Mean single sequence MFE | -20.63 |
| Consensus MFE | -10.21 |
| Energy contribution | -10.73 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.668872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 7941151 94 + 23011544 -------UUGUUGUUGCUGCCGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGACGACCCAGCGAACACCGAACACCCAACAACAACACCCA-- -------(((((((((((((((...)))))).(((((...............................))))).............)))))))))......-- ( -22.98, z-score = -3.15, R) >droSim1.chr2L 7736968 84 + 22036055 -----UGUUGUUGUUGCUGUCGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGACGACCCAGCGAACACCGAACACCCAA-------------- -----(((((.((((((((((((((.(((((...)))))..((...........))....)))))))..))).)))).).)))).....-------------- ( -22.20, z-score = -2.03, R) >droSec1.super_3 3455158 84 + 7220098 -----UGUUGUUGUUGCUGUCGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGACGACCCAGCGAACACCGAACACCCAA-------------- -----(((((.((((((((((((((.(((((...)))))..((...........))....)))))))..))).)))).).)))).....-------------- ( -22.20, z-score = -2.03, R) >droYak2.chr2L 17372117 82 - 22324452 -------UUGUUGUUGCUGUCGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGACGACCCAACGAACACCGAACACCCAA-------------- -------.((((((((..(((((((.(((((...)))))..((...........))....))))))).)))).))))............-------------- ( -20.80, z-score = -2.61, R) >droEre2.scaffold_4929 16863648 82 + 26641161 -------UUGUUGUUGCUGGCGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGACGACCCAACGAACACCCAACGCCACC-------------- -------.((((((((..(((((((.(((((...)))))..((...........))....))))).)))))).))))............-------------- ( -18.50, z-score = -0.76, R) >droWil1.scaffold_180703 569778 103 - 3946847 GUUGCCGUUGCCGUUGACGUCGUCUCAGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGUCGUCCUCUAGCCCAACUCCCACUUGCUGUUGCUGUUACUA (((..((....))..)))...((..(((((((((((.....((...........))....)).))))...((((.............)))).)))))..)).. ( -16.82, z-score = 1.07, R) >droGri2.scaffold_15252 4117843 76 - 17193109 --------UGGCUGCGUCGAUGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGUCAUGAGUUGACCCAGCCCCUCGCCCUCA------------------- --------.(((((.((((((((((....))))(((....)))..................)))))).)))))...........------------------- ( -20.90, z-score = -0.79, R) >consensus _______UUGUUGUUGCUGUCGUCUCGGCAGACGCUGCCCAGCCACCCAAUUAAGCCAUAAGACGACCCAGCGAACACCGAACACCCAA______________ .............((((((.(((((.(((((...)))))..((...........))....)))))...))))))............................. (-10.21 = -10.73 + 0.52)


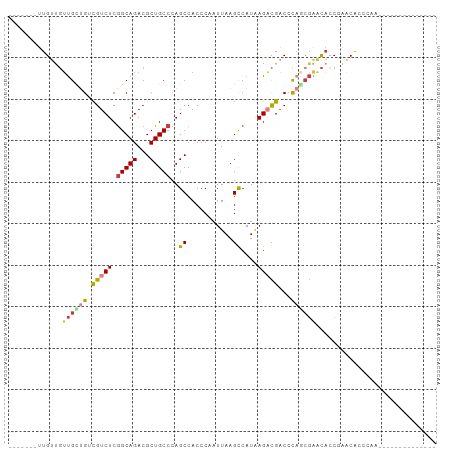
| Location | 7,941,151 – 7,941,245 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 73.52 |
| Shannon entropy | 0.49509 |
| G+C content | 0.56983 |
| Mean single sequence MFE | -30.66 |
| Consensus MFE | -17.29 |
| Energy contribution | -18.31 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.727522 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 7941151 94 - 23011544 --UGGGUGUUGUUGUUGGGUGUUCGGUGUUCGCUGGGUCGUCUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACGGCAGCAACAACAA------- --....(((((((((((....(((((((..(((((((((..((.((......)).))..))))...)))))..)))))))....))))))))))).------- ( -38.70, z-score = -3.11, R) >droSim1.chr2L 7736968 84 - 22036055 --------------UUGGGUGUUCGGUGUUCGCUGGGUCGUCUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACGACAGCAACAACAACA----- --------------.....((((...((((.(((..((((((((..((((.........)))).(((((...))))))))))))))))))))..))))----- ( -28.20, z-score = -0.68, R) >droSec1.super_3 3455158 84 - 7220098 --------------UUGGGUGUUCGGUGUUCGCUGGGUCGUCUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACGACAGCAACAACAACA----- --------------.....((((...((((.(((..((((((((..((((.........)))).(((((...))))))))))))))))))))..))))----- ( -28.20, z-score = -0.68, R) >droYak2.chr2L 17372117 82 + 22324452 --------------UUGGGUGUUCGGUGUUCGUUGGGUCGUCUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACGACAGCAACAACAA------- --------------............((((.((((.((((((((..((((.........)))).(((((...)))))))))))))..)))))))).------- ( -28.60, z-score = -1.47, R) >droEre2.scaffold_4929 16863648 82 - 26641161 --------------GGUGGCGUUGGGUGUUCGUUGGGUCGUCUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACGCCAGCAACAACAA------- --------------(.(((((((((.((..(((((((((((...))))))))).))..)).)).(((((...)))))..))))))).)........------- ( -30.60, z-score = -1.05, R) >droWil1.scaffold_180703 569778 103 + 3946847 UAGUAACAGCAACAGCAAGUGGGAGUUGGGCUAGAGGACGACUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCUGAGACGACGUCAACGGCAACGGCAAC ........((..((((.......(((((((((((((.....))).)))))))))).....))))(((..(((((....)))))..)))..((....))))... ( -31.00, z-score = -0.90, R) >droGri2.scaffold_15252 4117843 76 + 17193109 -------------------UGAGGGCGAGGGGCUGGGUCAACUCAUGACUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACAUCGACGCAGCCA-------- -------------------...........(((((.(((.........(......)((((.((.(((((...))))).)).))))))).))))).-------- ( -29.30, z-score = -1.17, R) >consensus ______________UUGGGUGUUCGGUGUUCGCUGGGUCGUCUUAUGGCUUAAUUGGGUGGCUGGGCAGCGUCUGCCGAGACGACAGCAACAACAA_______ ........................(.((((.((((...((((((..((((.........)))).(((((...))))))))))).)))))))).)......... (-17.29 = -18.31 + 1.03)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:24:46 2011