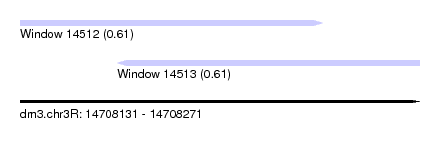

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 14,708,131 – 14,708,271 |
| Length | 140 |
| Max. P | 0.612429 |

| Location | 14,708,131 – 14,708,237 |
|---|---|
| Length | 106 |
| Sequences | 8 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 77.56 |
| Shannon entropy | 0.43048 |
| G+C content | 0.45196 |
| Mean single sequence MFE | -28.91 |
| Consensus MFE | -17.12 |
| Energy contribution | -16.23 |
| Covariance contribution | -0.89 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.607441 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 14708131 106 + 27905053 ACCUAGGCGAAAGUUUU-GCAUGG---AAGGAAUGUGUUUCCAGCACGUUUGCUCAAUGAGCAAACAACAAAUAGGGGAUUGGGGAUUCGGAUUCGGAUUUGGCUUCAGA------ .(((((((....))).(-((.(((---(((.......))))))))).((((((((...))))))))......))))...(((((((((((....)))))....)))))).------ ( -29.80, z-score = -1.28, R) >droSim1.chr3R 20749666 111 - 27517382 ACCUAGGCGAAAGUUUU-GCAUGGG-AGAGGAAUGUGUUGCCAGCACGUUUGCUCAAUGAGCAAACA---AAUAGGGGAUUGGGGAUUCGGAUUCGAAUACGGAUACGCCUUCAGA .(((((((((..((((.-.(.....-.)..))))...))))).....((((((((...)))))))).---..)))).((..((.((((((..........))))).).)).))... ( -32.50, z-score = -1.51, R) >droSec1.super_5 678446 111 - 5866729 ACCUAGGCGAAAGUUUU-GCAUGGU-AGAGGAAUGUGUUUCCAGCACGUUUGCUCAAUGAGCAAACA---AAUAGGGGAUUGGGGAUUCGGAUUCGGAUACGGAUUCGGCUUCAGA .((((.((....))(((-((.(.((-.(((((((((((.....)))))))).))).)).))))))..---..))))...((((((...((((((((....)))))))).)))))). ( -36.20, z-score = -2.82, R) >droYak2.chr3R 3371309 109 + 28832112 ACCUAGGCGAAAGUUUU-GCAUGG---GAGGUGUGUGUUUCCAGCACGUUUGCUCAAUGAGCAAACA---AAUAGGGGAUUGGGGAUCCGGAUUCGAAUUCGGAAUCGGGUACGGA .(((((((....))).(-((.(((---(((.......))))))))).((((((((...)))))))).---..))))...(((...((((((.((((....)))).)))))).))). ( -35.50, z-score = -1.93, R) >droEre2.scaffold_4770 10819877 109 - 17746568 ACCUAGGCGAAAGUUUU-GCAUGG---GAGGAAUGUGUUUCCAGCACGUUUGCUCAAUGAGCAAACA---AAUAGGGGAUUGGGGAUUCGGAUUCGGGUGCGGAUUGAAAACGGGA .((((.((((.....))-)).)))---).((((.....)))).((((((((((((...)))))))).---......(((((.(.....).)))))..))))............... ( -32.20, z-score = -1.79, R) >droAna3.scaffold_12911 3069843 109 + 5364042 ACCUAAGCGAAAGUUUU-GCAUGG--AAAUAUAUGUAUUUCGGGCACGUUUGUUCAAUGAGCAAACA---AAUAGGGGAUUGGGAAU-CGGACUUGGAUUCAGAUACGGAAUGGGA .(((((((....))).(-(((((.--.....))))))(((((.....((((((((...)))))))).---........(((..((((-(.......))))).))).))))))))). ( -25.60, z-score = -1.52, R) >dp4.chr2 14831125 107 + 30794189 ACCUAAACGAAAGUUUUUGCAUUGUAAGGGGAAAAUAUUUCGGGCACGUUUGCUCAAUGAGAAAGCA---AAUAGGGAAAUGGCAAAUGAGAAUGGGAUUGGGACUGGGA------ .(((((((....)))(((((..(((..(..(......)..)..))).(((((((.........))))---))).........)))))......)))).............------ ( -19.10, z-score = -0.06, R) >droPer1.super_0 6242392 107 - 11822988 ACCUAAACGAAAGUUUUUGCAUUGUAAGGGGAAAAUAUUUCGGGCAAGUUUGCUCAAUGAGAAAGCA---AAUAGGGAAAUGGCAAAUGAGAAUGGGACUGGGACUGGCA------ .(((((((....)))(((((.((((..(..(......)..)..))))(((((((.........))))---))).........)))))......)))).............------ ( -20.40, z-score = -0.07, R) >consensus ACCUAGGCGAAAGUUUU_GCAUGG__AGAGGAAUGUGUUUCCAGCACGUUUGCUCAAUGAGCAAACA___AAUAGGGGAUUGGGGAUUCGGAUUCGGAUUCGGAUUCGGA____GA .(((((((....)))...((.(((................)))))..((((((((...))))))))......))))........................................ (-17.12 = -16.23 + -0.89)


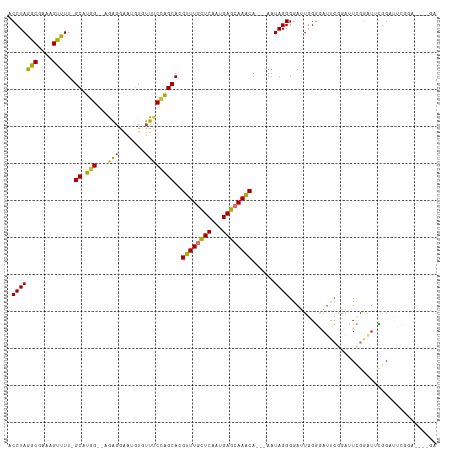
| Location | 14,708,165 – 14,708,271 |
|---|---|
| Length | 106 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 68.43 |
| Shannon entropy | 0.60300 |
| G+C content | 0.47740 |
| Mean single sequence MFE | -17.10 |
| Consensus MFE | -6.47 |
| Energy contribution | -7.10 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.612429 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
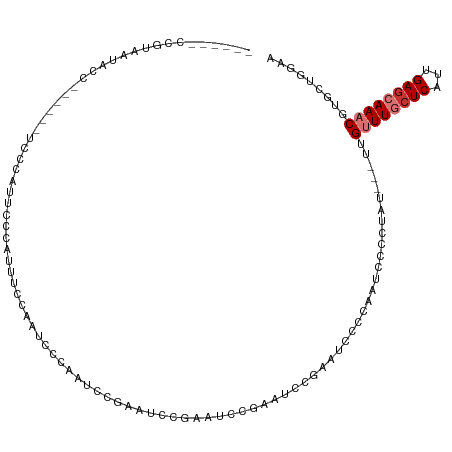
>dm3.chr3R 14708165 106 - 27905053 ACGUUGCCGUAAUACC------UCCCAUACCAAUUUUCAAUCUGAAGCCAAAUCC------GAAUCCGAAUCCCCAAUCCCCUAUUUGUUGUUUGCUCAUUGAGCAAACGUGCUGGAA (((....)))......------..((((((....((((.....))))........------.............................((((((((...))))))))))).))).. ( -14.60, z-score = -0.29, R) >droSim1.chr3R 20749702 109 + 27517382 ACGUUGCCGUAAUACC------UCCCAUGCCCAUUUUCAAUCUGAAGGCGUAUCCGUAUUCGAAUCCGAAUCCCCAAUCCCCUAU---UUGUUUGCUCAUUGAGCAAACGUGCUGGCA ....(((((((.....------....(((((....(((.....))))))))....(.(((((....))))).)............---..((((((((...)))))))).))).)))) ( -24.20, z-score = -2.12, R) >droSec1.super_5 678482 101 + 5866729 --------ACGUUACU------UCCCAUACCCAUUUUCAAUCUGAAGCCGAAUCCGUAUCCGAAUCCGAAUCCCCAAUCCCCUAU---UUGUUUGCUCAUUGAGCAAACGUGCUGGAA --------.((...((------((...................)))).))..(((((((.((....)).................---..((((((((...)))))))))))).))). ( -15.11, z-score = -0.90, R) >droYak2.chr3R 3371343 109 - 28832112 ACGUAGCCGUAAUACC------UCCCAUUCUAAUUUUUAAUCCGUACCCGAUUCCGAAUUCGAAUCCGGAUCCCCAAUCCCCUAU---UUGUUUGCUCAUUGAGCAAACGUGCUGGAA (((....)))......------..................((((((((((((((.(....))))).)))................---..((((((((...)))))))))))).))). ( -22.50, z-score = -2.40, R) >droEre2.scaffold_4770 10819911 115 + 17746568 GUGUAGCCGUAAUACCCCCAUUUCCCAUUCCCACUCCCACUCCCGUUUUCAAUCCGCACCCGAAUCCGAAUCCCCAAUCCCCUAU---UUGUUUGCUCAUUGAGCAAACGUGCUGGAA ....................................................(((((((.((....)).................---..((((((((...)))))))))))).))). ( -18.80, z-score = -2.60, R) >droAna3.scaffold_12911 3069878 102 - 5364042 ------GCGAAUUCCC------UGCCAAUCCCAAUACCAAUCCCAUUCCGUAUCUGAAUCCAAGUCCG-AUUCCCAAUCCCCUAU---UUGUUUGCUCAUUGAACAAACGUGCCCGAA ------(((.......------.................................(((((.......)-))))...........(---((((((.......))))))))))....... ( -8.10, z-score = 0.27, R) >dp4.chr2 14831163 99 - 30794189 -------------ACUC---GUACGGAUUCACGCUCCCAGUCCCAGUCCCAGUCCCAAUCCCAUUCUCAUUUGCCAUUUCCCUAU---UUGCUUUCUCAUUGAGCAAACGUGCCCGAA -------------....---(((((((((...((.....))...)))))...................................(---((((((.......))))))).))))..... ( -14.10, z-score = -1.21, R) >droPer1.super_0 6242430 99 + 11822988 -------------ACUC---GUACGGAUUCGCACUCCCAGUCCCAGUGCCAGUCCCAGUCCCAUUCUCAUUUGCCAUUUCCCUAU---UUGCUUUCUCAUUGAGCAAACUUGCCCGAA -------------..((---(..((((((.(((((.........))))).))))).............................(---((((((.......)))))))...)..))). ( -19.40, z-score = -2.66, R) >consensus ______CCGUAAUACC______UCCCAUUCCCAUUUCCAAUCCCAAUCCGAAUCCGAAUCCGAAUCCGAAUCCCCAAUCCCCUAU___UUGUUUGCUCAUUGAGCAAACGUGCUGGAA ..........................................................................................((((((((...))))))))......... ( -6.47 = -7.10 + 0.63)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:26:09 2011