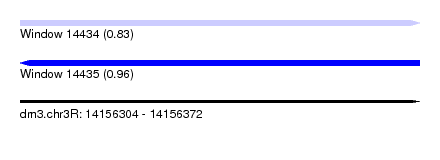

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 14,156,304 – 14,156,372 |
| Length | 68 |
| Max. P | 0.962398 |

| Location | 14,156,304 – 14,156,372 |
|---|---|
| Length | 68 |
| Sequences | 5 |
| Columns | 77 |
| Reading direction | forward |
| Mean pairwise identity | 68.11 |
| Shannon entropy | 0.55510 |
| G+C content | 0.52895 |
| Mean single sequence MFE | -18.02 |
| Consensus MFE | -10.64 |
| Energy contribution | -12.32 |
| Covariance contribution | 1.68 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.827533 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
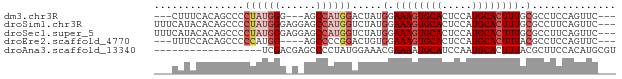
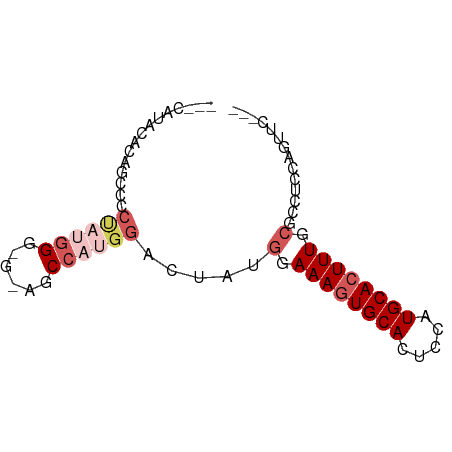
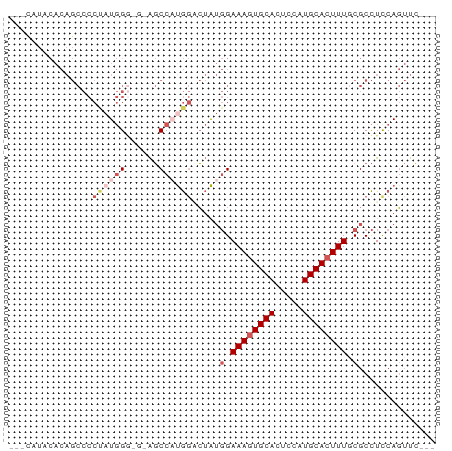
>dm3.chr3R 14156304 68 + 27905053 ---CUUUCACAGCCCCUAUGGG---AGCCAUGGACUAUGGAAAGUGCACUCCAUGCACUUUGCGCCUCCAGUUC--- ---........(((((...)))---.))..((((...((.((((((((.....)))))))).))..))))....--- ( -20.10, z-score = -1.18, R) >droSim1.chr3R 20197042 74 - 27517382 UUUCAUACACAGCCCCUAUGGGAGGAGCCAUGGUCUAUGGAAAGUGCACUCCAUGCACUUUGCGCCUUCAGUUC--- ....................(((((.((...(((.((((((........)))))).)))..)).))))).....--- ( -18.50, z-score = 0.07, R) >droSec1.super_5 125973 74 - 5866729 UUUCAUACACAGCCCCUAUGGGAGGAGCCAUGGUCUAUGGAAAGUGCACUCCAUGCACUUUGCGCCUUCAGUUC--- ....................(((((.((...(((.((((((........)))))).)))..)).))))).....--- ( -18.50, z-score = 0.07, R) >droEre2.scaffold_4770 10266990 67 - 17746568 ---UUUCCACAGCCCCCAUGG----AGCCCCGGACUGUGGAAAGUGCACUCCAUGCACUUUACGCCUCCAGUUC--- ---(((((((((.((....((----....)))).)))))))))(((((.....)))))................--- ( -20.50, z-score = -2.31, R) >droAna3.scaffold_13340 13754576 59 - 23697760 ------------------UCGACGAGCCCCUAUGGAAACGAAAAUGCAUCCAAUGCACUUUACGCUUCCACAUGCGU ------------------.......((.....(((((.(((((.(((((...))))).))).)).)))))...)).. ( -12.50, z-score = -1.81, R) >consensus ___CAUACACAGCCCCUAUGGG_G_AGCCAUGGACUAUGGAAAGUGCACUCCAUGCACUUUGCGCCUCCAGUUC___ ...............((((((......)))))).....(.((((((((.....)))))))).).............. (-10.64 = -12.32 + 1.68)
| Location | 14,156,304 – 14,156,372 |
|---|---|
| Length | 68 |
| Sequences | 5 |
| Columns | 77 |
| Reading direction | reverse |
| Mean pairwise identity | 68.11 |
| Shannon entropy | 0.55510 |
| G+C content | 0.52895 |
| Mean single sequence MFE | -24.80 |
| Consensus MFE | -11.68 |
| Energy contribution | -13.32 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.962398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
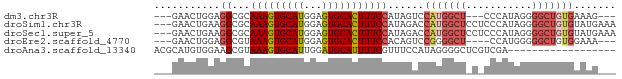
>dm3.chr3R 14156304 68 - 27905053 ---GAACUGGAGGCGCAAAGUGCAUGGAGUGCACUUUCCAUAGUCCAUGGCU---CCCAUAGGGGCUGUGAAAG--- ---..((((..((...(((((((((...))))))))))).)))).(((((((---((....)))))))))....--- ( -24.30, z-score = -1.51, R) >droSim1.chr3R 20197042 74 + 27517382 ---GAACUGAAGGCGCAAAGUGCAUGGAGUGCACUUUCCAUAGACCAUGGCUCCUCCCAUAGGGGCUGUGUAUGAAA ---........((...(((((((((...)))))))))))......((((((((((.....))))))))))....... ( -23.40, z-score = -0.93, R) >droSec1.super_5 125973 74 + 5866729 ---GAACUGAAGGCGCAAAGUGCAUGGAGUGCACUUUCCAUAGACCAUGGCUCCUCCCAUAGGGGCUGUGUAUGAAA ---........((...(((((((((...)))))))))))......((((((((((.....))))))))))....... ( -23.40, z-score = -0.93, R) >droEre2.scaffold_4770 10266990 67 + 17746568 ---GAACUGGAGGCGUAAAGUGCAUGGAGUGCACUUUCCACAGUCCGGGGCU----CCAUGGGGGCUGUGGAAA--- ---...(((..((...(((((((((...))))))))))).)))((((.((((----((...)))))).))))..--- ( -27.70, z-score = -2.25, R) >droAna3.scaffold_13340 13754576 59 + 23697760 ACGCAUGUGGAAGCGUAAAGUGCAUUGGAUGCAUUUUCGUUUCCAUAGGGGCUCGUCGA------------------ ..((.((((((((((.(((((((((...)))))))))))))))))))...)).......------------------ ( -25.20, z-score = -4.29, R) >consensus ___GAACUGGAGGCGCAAAGUGCAUGGAGUGCACUUUCCAUAGACCAUGGCU_C_CCCAUAGGGGCUGUGUAAG___ ..............(.(((((((((...))))))))))(((((.((((((......))))..)).)))))....... (-11.68 = -13.32 + 1.64)
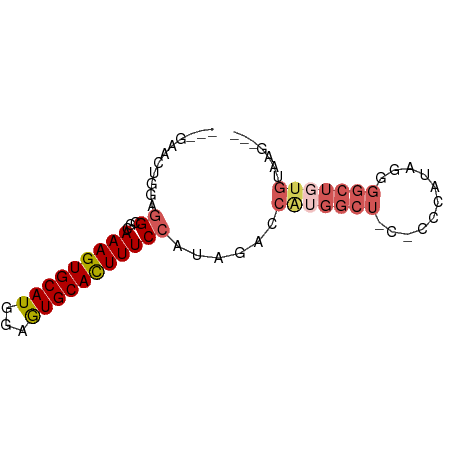
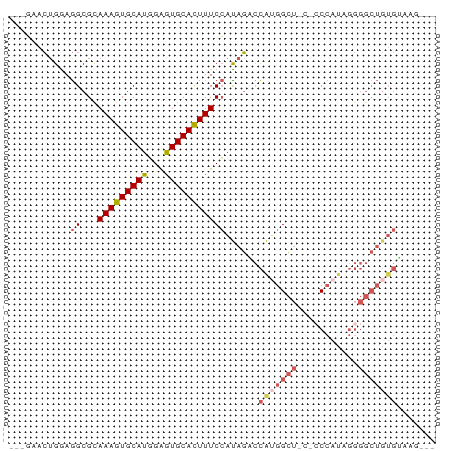
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:25:03 2011