
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,781,915 – 13,782,009 |
| Length | 94 |
| Max. P | 0.991928 |

| Location | 13,781,915 – 13,782,009 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 93.76 |
| Shannon entropy | 0.10514 |
| G+C content | 0.51592 |
| Mean single sequence MFE | -30.60 |
| Consensus MFE | -24.72 |
| Energy contribution | -24.88 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.624695 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
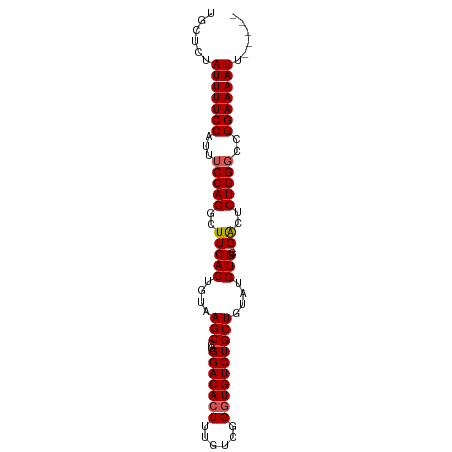
>dm3.chr3R 13781915 94 + 27905053 UGCUCUAUUUUCCAUUUCCAGGCUUCACUGUAAGCCGGGACACUUUGUCGGCUGUCUGCUGUAUGUGUGUGACUCUGGGCCGGAAAAUUUGGGC .((((.(((((((...(((((((((......)))))(..((((.....((((.....)))).....))))..)..))))..)))))))..)))) ( -31.10, z-score = -1.81, R) >droEre2.scaffold_4770 9901176 87 - 17746568 UGCCCUAUUUUCCAGUUCCAGGCUUCACUGUAAGCCGGGACACUUUGUCGGGUGUCUGCUGUAUGUGC--GGCUCUGGGCCGGAAAAUU----- ......(((((((.(..(((((((.(((....(((..(((((((......))))))))))....))).--))).))))..)))))))).----- ( -34.40, z-score = -2.52, R) >droYak2.chr3R 2374431 87 + 28832112 UGCUCUAUUUUCCAUUGCCAGGCUUCACUGUAAGCCGGGACACUUUGUCGGGUGUCUGCUGUAUGUGC--GGCUCUGGGCCGGAAAAUU----- ......(((((((....(((((((.(((....(((..(((((((......))))))))))....))).--))).))))...))))))).----- ( -31.60, z-score = -1.96, R) >droSec1.super_12 1749305 87 - 2123299 UGCUCUAUUUUCCAUUUCCAGGCUUCACUGUAAGCCGGGACACUUUGUCGGGUGUCUGCUGUAUGUGC--GACUCUGGGCCGGAAAAUU----- ......(((((((...(((((..(((((....(((..(((((((......))))))))))....))).--))..)))))..))))))).----- ( -27.80, z-score = -1.60, R) >droSim1.chr3R 19808208 87 - 27517382 UGCUCUAUUUUCCAUUUCCAGGCUUCACUGUAAGCCGGGACACUUUGUCGGGUGUCUGCUGUAUGUGU--GACUCUGGGCCGGAAAAUU----- ......(((((((...((((((..(((((((((((..(((((((......)))))))))).)))).))--)).))))))..))))))).----- ( -28.10, z-score = -1.86, R) >consensus UGCUCUAUUUUCCAUUUCCAGGCUUCACUGUAAGCCGGGACACUUUGUCGGGUGUCUGCUGUAUGUGC__GACUCUGGGCCGGAAAAUU_____ ......(((((((...(((((..(((((....(((..(((((((......))))))))))....)))...))..)))))..)))))))...... (-24.72 = -24.88 + 0.16)


| Location | 13,781,915 – 13,782,009 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 93.76 |
| Shannon entropy | 0.10514 |
| G+C content | 0.51592 |
| Mean single sequence MFE | -27.72 |
| Consensus MFE | -25.28 |
| Energy contribution | -25.24 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.92 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.51 |
| SVM RNA-class probability | 0.991928 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
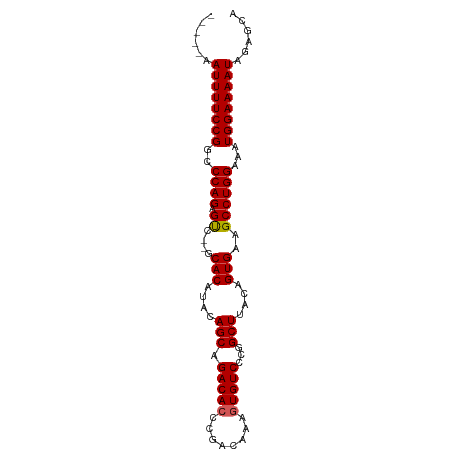
>dm3.chr3R 13781915 94 - 27905053 GCCCAAAUUUUCCGGCCCAGAGUCACACACAUACAGCAGACAGCCGACAAAGUGUCCCGGCUUACAGUGAAGCCUGGAAAUGGAAAAUAGAGCA ((....((((((((..((((..((((.......(....)..(((((((.....)))..))))....))))...))))...))))))))...)). ( -24.50, z-score = -1.86, R) >droEre2.scaffold_4770 9901176 87 + 17746568 -----AAUUUUCCGGCCCAGAGCC--GCACAUACAGCAGACACCCGACAAAGUGUCCCGGCUUACAGUGAAGCCUGGAACUGGAAAAUAGGGCA -----.(((((((((.((((.((.--.(((....(((.(((((........)))))...)))....)))..))))))..)))))))))...... ( -30.40, z-score = -2.80, R) >droYak2.chr3R 2374431 87 - 28832112 -----AAUUUUCCGGCCCAGAGCC--GCACAUACAGCAGACACCCGACAAAGUGUCCCGGCUUACAGUGAAGCCUGGCAAUGGAAAAUAGAGCA -----.((((((((..((((.((.--.(((....(((.(((((........)))))...)))....)))..))))))...))))))))...... ( -28.70, z-score = -3.25, R) >droSec1.super_12 1749305 87 + 2123299 -----AAUUUUCCGGCCCAGAGUC--GCACAUACAGCAGACACCCGACAAAGUGUCCCGGCUUACAGUGAAGCCUGGAAAUGGAAAAUAGAGCA -----.((((((((..((((.((.--.(((....(((.(((((........)))))...)))....)))..))))))...))))))))...... ( -28.20, z-score = -3.40, R) >droSim1.chr3R 19808208 87 + 27517382 -----AAUUUUCCGGCCCAGAGUC--ACACAUACAGCAGACACCCGACAAAGUGUCCCGGCUUACAGUGAAGCCUGGAAAUGGAAAAUAGAGCA -----.((((((((..((((.((.--.(((....(((.(((((........)))))...)))....)))..))))))...))))))))...... ( -26.80, z-score = -3.30, R) >consensus _____AAUUUUCCGGCCCAGAGUC__GCACAUACAGCAGACACCCGACAAAGUGUCCCGGCUUACAGUGAAGCCUGGAAAUGGAAAAUAGAGCA ......((((((((..((((.((....(((....(((.(((((........)))))...)))....)))..))))))...))))))))...... (-25.28 = -25.24 + -0.04)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:24:00 2011