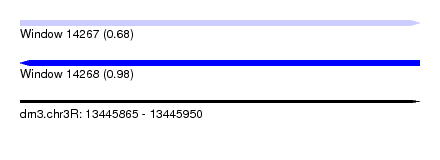

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,445,865 – 13,445,950 |
| Length | 85 |
| Max. P | 0.979574 |

| Location | 13,445,865 – 13,445,950 |
|---|---|
| Length | 85 |
| Sequences | 13 |
| Columns | 85 |
| Reading direction | forward |
| Mean pairwise identity | 88.24 |
| Shannon entropy | 0.28917 |
| G+C content | 0.56876 |
| Mean single sequence MFE | -24.04 |
| Consensus MFE | -22.58 |
| Energy contribution | -22.62 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.17 |
| Mean z-score | -0.80 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.682119 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
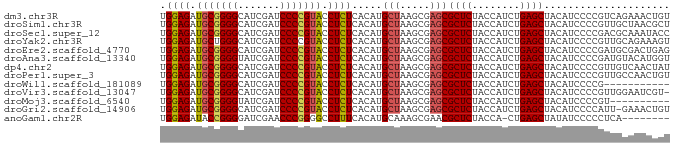
>dm3.chr3R 13445865 85 + 27905053 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUCAGAAACUGU .((((.(((((((.......))))))).)))).....(((.....)))((((........))))..................... ( -23.70, z-score = -0.47, R) >droSim1.chr3R 19485020 85 - 27517382 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUUGCUAACGCU .((((.(((((((.......))))))).)))).........(((((((((((........)))).((......)).)))..)))) ( -26.60, z-score = -1.27, R) >droSec1.super_12 1407740 85 - 2123299 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGACGCAAAUACC .((((.(((((((.......))))))).))))..........(((...((((........))))..(......).)))....... ( -23.90, z-score = -1.14, R) >droYak2.chr3R 2006652 85 + 28832112 UGGAGAUGCUGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUUGCAGAAAGU .((.((((..(((........)))......((((.(((.(((((....))).)).))).))))...)))).))............ ( -17.80, z-score = 1.29, R) >droEre2.scaffold_4770 9560487 85 - 17746568 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGAUGCGACUGAG ..(((.(((((((.......))))))).)))((.((((((.....)))((((........))))..........))).))..... ( -25.10, z-score = -0.56, R) >droAna3.scaffold_13340 5736719 85 + 23697760 UGGAGAUGCGGGGUAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGAUGUACAUGGU .((((.(((((((.......))))))).))))..((((..........((((........))))((((((...)))))))))).. ( -29.40, z-score = -1.78, R) >dp4.chr2 7041878 85 - 30794189 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUUGUCAACUAU .((((.(((((((.......))))))).)))).....(((.....)))((((........))))..................... ( -23.70, z-score = -0.81, R) >droPer1.super_3 845234 85 - 7375914 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUUGCCAACUGU .((((.(((((((.......))))))).)))).....(((.....)))((((........))))..................... ( -23.70, z-score = -0.36, R) >droWil1.scaffold_181089 3825657 74 - 12369635 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCG----------- .((((.(((((((.......))))))).)))).....(((.....)))((((........))))..........----------- ( -23.70, z-score = -1.57, R) >droVir3.scaffold_13047 2062971 84 - 19223366 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUUGGAAUCGU- .((((.(((((((.......))))))).))))..........((((..((((........))))....(((.....))).))))- ( -26.10, z-score = -0.73, R) >droMoj3.scaffold_6540 19611495 75 - 34148556 UGGAGAUGCGGGGUAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGU---------- .((((.(((((((.......))))))).)))).....(((.....)))((((........))))...........---------- ( -24.70, z-score = -1.60, R) >droGri2.scaffold_14906 1941405 84 - 14172833 UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCAUU-GAAACUGU .((((.(((((((.......))))))).)))).....(((.....)))((((........))))............-........ ( -23.70, z-score = -1.04, R) >anoGam1.chr2R 51279388 76 - 62725911 UGGAGAUACCGGGGAUCGAACCCGGGGCCUUUCACAUGCAAAGCGAACGCUCUACCA-CUGAGCUAUAUCCCCCUCA-------- .((.(((((((((.......))))).......................((((.....-..))))..)))).))....-------- ( -20.40, z-score = -0.36, R) >consensus UGGAGAUGCGGGGCAUCGAUCCCCGUACCUCUCACAUGCUAAGCGAGCGCUCUACCAUCUGAGCUACAUCCCCGUUGCAAACGGU .((((.(((((((.......))))))).)))).....(((.....)))((((........))))..................... (-22.58 = -22.62 + 0.04)

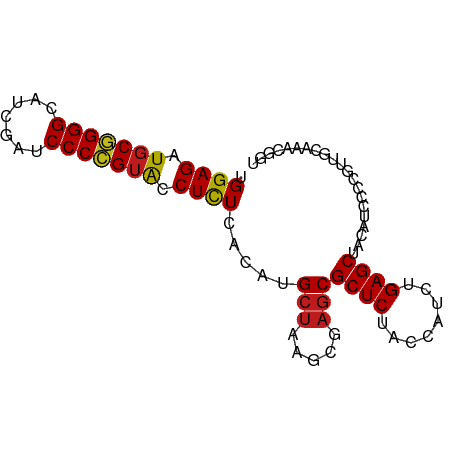
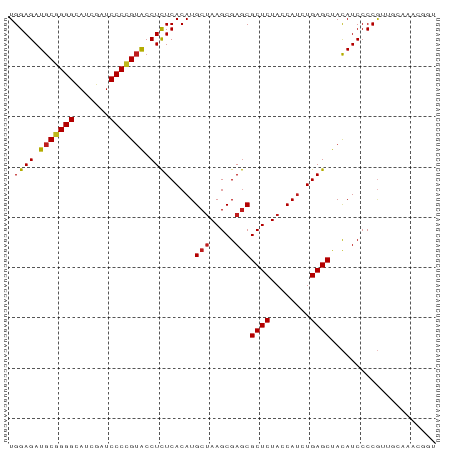
| Location | 13,445,865 – 13,445,950 |
|---|---|
| Length | 85 |
| Sequences | 13 |
| Columns | 85 |
| Reading direction | reverse |
| Mean pairwise identity | 88.24 |
| Shannon entropy | 0.28917 |
| G+C content | 0.56876 |
| Mean single sequence MFE | -32.70 |
| Consensus MFE | -31.49 |
| Energy contribution | -31.53 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.96 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.979574 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
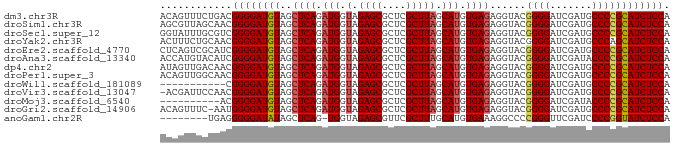
>dm3.chr3R 13445865 85 - 27905053 ACAGUUUCUGACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA .(((...)))..((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -33.20, z-score = -1.62, R) >droSim1.chr3R 19485020 85 + 27517382 AGCGUUAGCAACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA .((....))...((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -35.00, z-score = -2.09, R) >droSec1.super_12 1407740 85 + 2123299 GGUAUUUGCGUCGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA ............((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -33.10, z-score = -1.30, R) >droYak2.chr3R 2006652 85 - 28832112 ACUUUCUGCAACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCAGCAUCUCCA ............((((((((..((((.(((.(.((((....))))).))).))))......(((........))).)))))))). ( -30.20, z-score = -0.92, R) >droEre2.scaffold_4770 9560487 85 + 17746568 CUCAGUCGCAUCGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA ............((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -33.10, z-score = -1.08, R) >droAna3.scaffold_13340 5736719 85 - 23697760 ACCAUGUACAUCGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUACCCCGCAUCUCCA ............((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -32.90, z-score = -1.86, R) >dp4.chr2 7041878 85 + 30794189 AUAGUUGACAACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA ............((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -33.10, z-score = -2.04, R) >droPer1.super_3 845234 85 + 7375914 ACAGUUGGCAACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA ......(....)((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -34.10, z-score = -1.85, R) >droWil1.scaffold_181089 3825657 74 + 12369635 -----------CGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA -----------.((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -33.10, z-score = -2.57, R) >droVir3.scaffold_13047 2062971 84 + 19223366 -ACGAUUCCAACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA -...........((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -33.10, z-score = -1.77, R) >droMoj3.scaffold_6540 19611495 75 + 34148556 ----------ACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUACCCCGCAUCUCCA ----------..((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). ( -32.90, z-score = -2.86, R) >droGri2.scaffold_14906 1941405 84 + 14172833 ACAGUUUC-AAUGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA ........-..(((((((((..((((.(((.(.((((....))))).))).))))......((((.......))))))))))))) ( -33.00, z-score = -2.10, R) >anoGam1.chr2R 51279388 76 + 62725911 --------UGAGGGGGAUAUAGCUCAG-UGGUAGAGCGUUCGCUUUGCAUGUGAAAGGCCCCGGGUUCGAUCCCCGGUAUCUCCA --------....(((((((..(((((.-..(((((((....)))))))...)))...)).(((((.......)))))))))))). ( -28.30, z-score = -1.23, R) >consensus ACAGUUUGCAACGGGGAUGUAGCUCAGAUGGUAGAGCGCUCGCUUAGCAUGUGAGAGGUACGGGGAUCGAUGCCCCGCAUCUCCA ............((((((((..((((.(((.(.((((....))))).))).))))......((((.......)))))))))))). (-31.49 = -31.53 + 0.04)

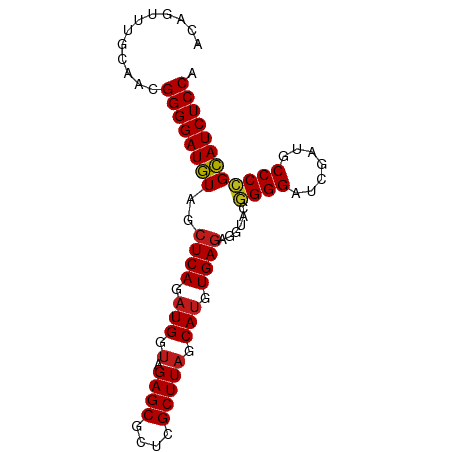
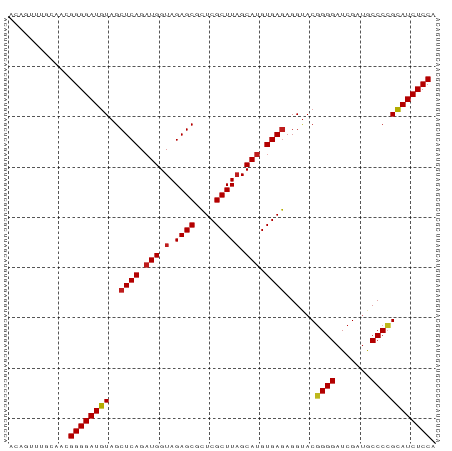
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:22:46 2011