
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,401,960 – 13,402,041 |
| Length | 81 |
| Max. P | 0.871532 |

| Location | 13,401,960 – 13,402,041 |
|---|---|
| Length | 81 |
| Sequences | 3 |
| Columns | 84 |
| Reading direction | reverse |
| Mean pairwise identity | 45.38 |
| Shannon entropy | 0.79773 |
| G+C content | 0.52505 |
| Mean single sequence MFE | -23.27 |
| Consensus MFE | -9.27 |
| Energy contribution | -9.17 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.871532 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
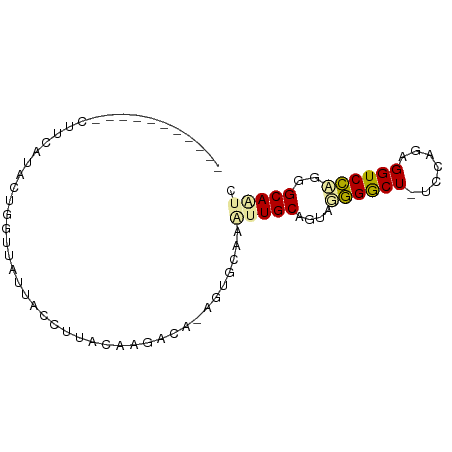
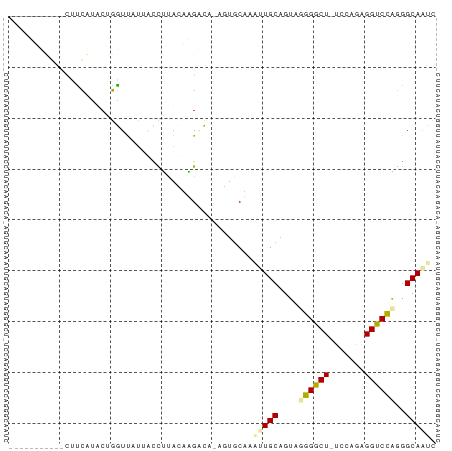
>dm3.chr3R 13401960 81 - 27905053 --UCCUUUAUCCUUUAUCCUAAUCCUACCCUCGCAGGAUGCCGUGCUAAUUGCUGUUGCGGCU-UACAAUGGUCGAGGGCAGUC --......................((.(((((((((.(.((.((....)).))).))))((((-......))))))))).)).. ( -20.20, z-score = -0.44, R) >droAna3.scaffold_13250 1227727 70 + 3535662 ------------UCCUUUCUGGUUUUUAUUAUAUUUGCCU-UCUGCACGUUGCAGAAGGGACUCUCC-GAGGUCCUGGGCAAUC ------------.((.....))............((((((-(((((.....))))))((((((....-..)))))).))))).. ( -21.90, z-score = -2.29, R) >droVir3.scaffold_12736 587294 82 - 612119 CCGGUGCCGGUCUUCGCACAGGUUAUUGCCUUACCAGGCA-AGU-CGAUCUGCAGUGUGGGCUGUCCAGAGGUCCAGGGCAAUC ....((((((.((((...((((((.((((((....)))))-)..-.))))))((((....))))....)))).))..))))... ( -27.70, z-score = -0.37, R) >consensus ___________CUUCAUACUGGUUAUUACCUUACAAGACA_AGUGCAAAUUGCAGUAGGGGCU_UCCAGAGGUCCAGGGCAAUC ................................................(((((....((((((.......))))))..))))). ( -9.27 = -9.17 + -0.10)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:22:38 2011