
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,262,227 – 13,262,381 |
| Length | 154 |
| Max. P | 0.569041 |

| Location | 13,262,227 – 13,262,381 |
|---|---|
| Length | 154 |
| Sequences | 6 |
| Columns | 167 |
| Reading direction | forward |
| Mean pairwise identity | 78.86 |
| Shannon entropy | 0.39195 |
| G+C content | 0.45433 |
| Mean single sequence MFE | -47.41 |
| Consensus MFE | -31.59 |
| Energy contribution | -31.60 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.569041 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
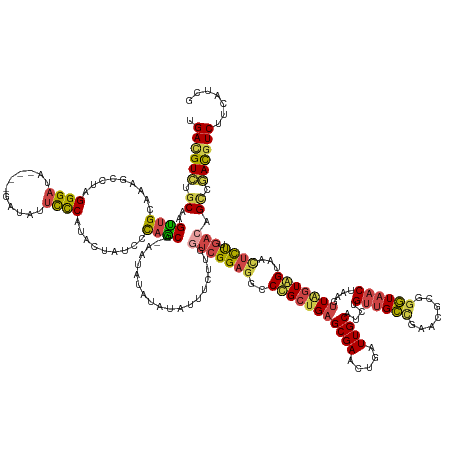
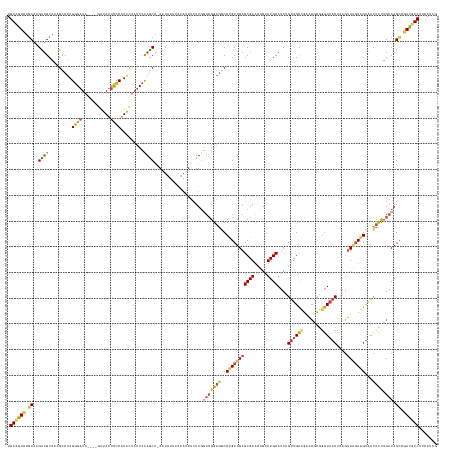
>dm3.chr3R 13262227 154 + 27905053 UGACGUCUGCAAGUUGCACAGCCUAGGGAUA----GAUAUUCCCAUACCAUCCCAGCA-AAUAU--------CUUGGUCGGAGGCCUGCUGAGCGAACUGAUUGCAUCUGUUGCUGAACGCGGGUAACUAAGUUAGUAGUAACUUUUGACAGCCGACGUCUUCAUCG .((((((.((.((((((...((((.((((..----.....))))...((..((.((..-.....--------)).))..))))))(((((.(((((...(((...)))..))))).)..))))))))))((((((....))))))......)).))))))....... ( -46.80, z-score = -1.11, R) >droSim1.chr3R 19309061 162 - 27517382 UGACGUCUGCAAGUUGCACAGCCUAGGGAUA----AAUAUUCCCAUACUAUCCCAGCA-AAUAUAUAUAUUUCUUGGUCGGAGGCCUGCUGAGCGAACUGAUUGCAUCUGUUGCUGAACGCGGGUAACUAAGUUAGUAGUAACUCCUGACAGCCGACGUCUUCAUCG .((((((.((.((((((...((((.((((..----.....))))...((..((.((.(-((((....))))))).))..))))))(((((.(((((...(((...)))..))))).)..))))))))))..(((((.((...)).))))).)).))))))....... ( -49.40, z-score = -2.00, R) >droSec1.super_12 1239976 162 - 2123299 UGACGUCUGCAUGUUGCACAGUCUAGGGAUA----AAUAUUCCCAUAUUAUCCCAGCA-AAUAUAUAUAUUUCUUGGUCGGAGGCCUGCUGAGCGAACUGAUUGCAUCUGUUGCUGAACGCGGGUAACUAAGUUAGUAGUGACUCCUGACAGCCGACGUCUUUAUCG .((((((.((.(((((.........((((..----.....)))).........)))))-.................((((((((((((((.(((((...(((...)))..))))).)..)))))).((((......))))..)))).))).)).))))))....... ( -47.67, z-score = -1.39, R) >droYak2.chr3R 1810538 162 + 28832112 UGACGUCUGCAAGUUGCAAUGCCUAGGGAUA----GAUAUUCCCAUACUAUCCUAACA-AAUAUAUAUAUUUCUUGUUCGGAAGCCCGCUGAGCGAACUGAUUGCAUCUGUUGCCGCUCACGGGUAACUGAGUUAGUGGUAGCUCUUGACAGUCAAUAUCAUCAUUA ....(.(((((((..((........((((..----.....))))....((((((((((-((((....))))).....((((..(((((.(((((((((.(((...))).)))..)))))))))))..))))))))).)))))).)))).))).)............. ( -47.20, z-score = -1.90, R) >droEre2.scaffold_4770 9383951 162 - 17746568 UGACGUCUGCAAGCUGCAAUGCCUAGGGAUA----GAUAUUCCCAUACUAUCCUAGCA-AAUAUAUAUAUUUCUUGGUAGGAGGCCCGCUGAGCGAACUGAUUGCAUCUGUUGCCGAGCACGGGUAACUAAGUUGGUGGUAACUCUUGACAGCCGACGUCUUCAUUG .((((((.....(((((((.(((..((((..----.....))))......(((((.((-(.............))).))))))))((((..(((((.....))))....((((((.......))))))....)..))))......))).)))).))))))....... ( -49.42, z-score = -1.15, R) >droAna3.scaffold_13340 5817833 165 - 23697760 UGAUGUUUUCAUGCGGCAAUGCCCAGGGAUAUAGAGGUACAUUCGUGCAUACAUAGCUCAGUAUAUAUAUUUUUUUUGCGGCGGCCCGCUGAGCGAUGAGGUUGCACCAGA-GUCGCUACUAUAUG-CUAAUAUUUUAGCAAUCUUUGACAGCCAAUGUCUUCAUUG .((((....(((..(((..((..(((((((((((.(((..(((((((((..(((.((((((((....))).......((((....))))))))).)))....)))))..))-)).))).)))))((-((((....)))))).)))))).))))).)))....)))). ( -44.00, z-score = 0.11, R) >consensus UGACGUCUGCAAGUUGCAAAGCCUAGGGAUA____GAUAUUCCCAUACUAUCCCAGCA_AAUAUAUAUAUUUCUUGGUCGGAGGCCCGCUGAGCGAACUGAUUGCAUCUGUUGCCGAACGCGGGUAACUAAGUUAGUAGUAACUCUUGACAGCCGACGUCUUCAUCG .((((((.((..((((.........((((...........)))).........))))...................(((((((..(((((((((((.....))))....((((((.......))))))....)))))))...)))).))).)).))))))....... (-31.59 = -31.60 + 0.01)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:22:28 2011