
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,259,976 – 13,260,073 |
| Length | 97 |
| Max. P | 0.946714 |

| Location | 13,259,976 – 13,260,073 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 71.28 |
| Shannon entropy | 0.52421 |
| G+C content | 0.52671 |
| Mean single sequence MFE | -22.55 |
| Consensus MFE | -14.19 |
| Energy contribution | -14.58 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.946714 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
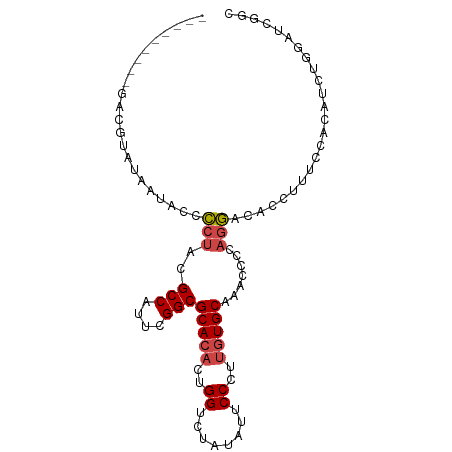
>dm3.chr3R 13259976 97 + 27905053 GACGUGUUGUACCCCUAAUAUGCCUACGCCAUUCGGCGCACACUGGUCUAUAUUCCCUUGUGCACGGCGAAGGACACUGUUCCACAAUUGGAUCGGC ..(((((((......)))))))(((.((((.....(.(((((..((........))..))))).))))).)))...(((.((((....)))).))). ( -27.00, z-score = -0.49, R) >droSim1.chr3R 19306757 88 - 27517382 ---------GACGUAUUAUACCCCUACGCCAUUCGGCGCACACUGGUCUAUAUUCCCUUGUGCAAACCCCAGGACACCUUUCCACACCUGGAUCGGC ---------.............(((..(((....)))(((((..((........))..))))).......)))...((..((((....))))..)). ( -22.10, z-score = -1.99, R) >droSec1.super_12 1237687 88 - 2123299 ---------GACGUAUUAUACCCCUACGCCAUUCGGCGCACACUGGUCUAUAUUCCCUUGUGCAAACCCCAGGACACCUUUCCACACCUGGAUCGGC ---------.............(((..(((....)))(((((..((........))..))))).......)))...((..((((....))))..)). ( -22.10, z-score = -1.99, R) >droYak2.chr3R 1808290 88 + 28832112 ---------GACGUACUAUGCCCCUACGCCAUUCGGCGCACACUGGACUAUAUUCCCUUGUGCAAACCUGAGGACACCUCACCAAAUCUGAAUCGGC ---------..................(((((((((.(((((..(((......)))..))))).....(((((...)))))......)))))).))) ( -23.90, z-score = -2.99, R) >droEre2.scaffold_4770 9381633 88 - 17746568 ---------GACGUAUAAUACCCCUACGCCAUUCGGCGCACACUGGUCUAUACUCCCUUGUGCAAGCACGAGGACAUCUCUCCGGAUCUGAAUUGGC ---------..................(((((((((......((((.........(((((((....)))))))........))))..))))).)))) ( -22.03, z-score = -0.86, R) >droWil1.scaffold_181089 1859988 86 + 12369635 ---------GCCAUUGAUUAGGACGACGCCAGAUGGCGCAAACAGGACGGAAUUCCCGAGUGCAUGGUAUAUUACAACGAAU--AUUUGGGAUUGAU ---------((((((.....((......)).))))))................(((((((((..((...........))..)--))))))))..... ( -18.20, z-score = -0.19, R) >consensus _________GACGUAUAAUACCCCUACGCCAUUCGGCGCACACUGGUCUAUAUUCCCUUGUGCAAACCCCAGGACACCUUUCCACAUCUGGAUCGGC ......................(((..(((....)))(((((..((........))..))))).......)))........................ (-14.19 = -14.58 + 0.39)


| Location | 13,259,976 – 13,260,073 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 71.28 |
| Shannon entropy | 0.52421 |
| G+C content | 0.52671 |
| Mean single sequence MFE | -26.48 |
| Consensus MFE | -17.48 |
| Energy contribution | -17.40 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.29 |
| Mean z-score | -0.92 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.806872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
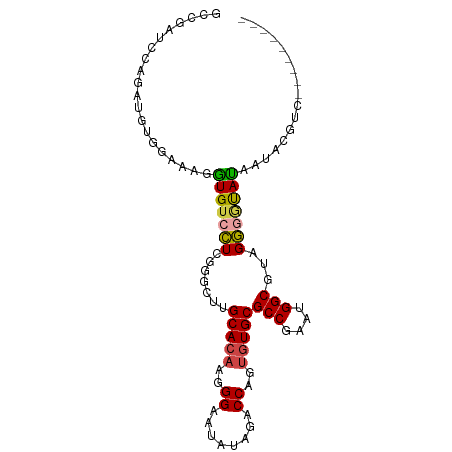
>dm3.chr3R 13259976 97 - 27905053 GCCGAUCCAAUUGUGGAACAGUGUCCUUCGCCGUGCACAAGGGAAUAUAGACCAGUGUGCGCCGAAUGGCGUAGGCAUAUUAGGGGUACAACACGUC ((((.((((....)))).).(((.(((.(((((((((((..((........))..))))))).....)))).))))))......))).......... ( -32.20, z-score = -1.18, R) >droSim1.chr3R 19306757 88 + 27517382 GCCGAUCCAGGUGUGGAAAGGUGUCCUGGGGUUUGCACAAGGGAAUAUAGACCAGUGUGCGCCGAAUGGCGUAGGGGUAUAAUACGUC--------- (((..((((....))))..))).(((((......(((((..((........))..)))))(((....))).)))))............--------- ( -26.40, z-score = -0.54, R) >droSec1.super_12 1237687 88 + 2123299 GCCGAUCCAGGUGUGGAAAGGUGUCCUGGGGUUUGCACAAGGGAAUAUAGACCAGUGUGCGCCGAAUGGCGUAGGGGUAUAAUACGUC--------- (((..((((....))))..))).(((((......(((((..((........))..)))))(((....))).)))))............--------- ( -26.40, z-score = -0.54, R) >droYak2.chr3R 1808290 88 - 28832112 GCCGAUUCAGAUUUGGUGAGGUGUCCUCAGGUUUGCACAAGGGAAUAUAGUCCAGUGUGCGCCGAAUGGCGUAGGGGCAUAGUACGUC--------- (((((.......)))))...(((((((.......(((((..(((......)))..)))))(((....)))...)))))))........--------- ( -29.20, z-score = -1.68, R) >droEre2.scaffold_4770 9381633 88 + 17746568 GCCAAUUCAGAUCCGGAGAGAUGUCCUCGUGCUUGCACAAGGGAGUAUAGACCAGUGUGCGCCGAAUGGCGUAGGGGUAUUAUACGUC--------- ...........(((...(((.....)))(((....)))...)))(((((((((....((((((....))))))..))).))))))...--------- ( -25.70, z-score = -0.36, R) >droWil1.scaffold_181089 1859988 86 - 12369635 AUCAAUCCCAAAU--AUUCGUUGUAAUAUACCAUGCACUCGGGAAUUCCGUCCUGUUUGCGCCAUCUGGCGUCGUCCUAAUCAAUGGC--------- .............--..((((((...........(((..(((((......)))))..)))(((....)))...........)))))).--------- ( -19.00, z-score = -1.24, R) >consensus GCCGAUCCAGAUGUGGAAAGGUGUCCUCGGGCUUGCACAAGGGAAUAUAGACCAGUGUGCGCCGAAUGGCGUAGGGGUAUAAUACGUC_________ ....................(((((((.......(((((..((........))..)))))(((....)))...)))))))................. (-17.48 = -17.40 + -0.08)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:22:26 2011