
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,207,243 – 13,207,342 |
| Length | 99 |
| Max. P | 0.859296 |

| Location | 13,207,243 – 13,207,342 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 66.48 |
| Shannon entropy | 0.65002 |
| G+C content | 0.43067 |
| Mean single sequence MFE | -19.61 |
| Consensus MFE | -6.22 |
| Energy contribution | -6.73 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.859296 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 13207243 99 + 27905053 GUCUAUAAAAUUAUCCGCAAAUUAAAGCGUCUAAAGCCACAGACGCUGGCUGAAAAGCUGAAAA-GCUGCAACGCUGAAAGACUGAAAAACCAAAAGACA-- ((((.....................(((((((........)))))))(((.....((((....)-))).....)))...................)))).-- ( -22.80, z-score = -2.52, R) >droSim1.chr3R 19252237 99 - 27517382 GUCUAUAAAAUUAUCCGCAAAUUAAAGCGUCUAAAGCCAAAGACGCUGGCUGCAAAGCUGAAAA-GCUGCAACGCUGAAAGACUGAAAAACCAAAAGACA-- ((((.....................(((((((........)))))))(((((((..(((....)-))))))..)))...................)))).-- ( -24.00, z-score = -2.58, R) >droSec1.super_12 1182102 99 - 2123299 GUCUAUAAAAUUAUCCGCAAAUUAAAGCGUCUAAAGCCAAAGACGCUGGCUGCAAAGCUGAAAA-GCUGCAACGCUGAAAGACUGAAAAACCAAAAGACA-- ((((.....................(((((((........)))))))(((((((..(((....)-))))))..)))...................)))).-- ( -24.00, z-score = -2.58, R) >droEre2.scaffold_4770 9324023 90 - 17746568 GUCUAUAAAAUUAUCCGCAAAUUAAAGCGUCUAA-GCCAAAGACGCUGGCUGAAAAACUGAAAA------GCCGCAGAGC---UGAAAAACCAAAAGACA-- ((((.........((.((.......(((((((..-.....)))))))((((............)------))).....))---.)).........)))).-- ( -21.47, z-score = -3.16, R) >droAna3.scaffold_13340 10418445 83 + 23697760 GUCUAUAAAAUUAUCUGUAAAUUAAAGCGUCUAAGCCCA-AGACGCCAAGACGAGUGGCUCACG------GGCUGAUAGUC--CGAAAGACC---------- ((((.....((((((.((........)).....(((((.-.((.((((.......))))))..)------)))))))))).--....)))).---------- ( -21.00, z-score = -1.91, R) >dp4.chr2 21573938 80 - 30794189 GUCUAUAAAAUUAUCUGCAAAUUAAAGCGUCUAAAGGCGCAGACGCCAGGCCGAC--CCAAAGA----------CAGAGCCCCCAACCGACA---------- (((..........(((((........))((((...((((....)))).((.....--))..)))----------))))..........))).---------- ( -16.25, z-score = -1.69, R) >droPer1.super_0 8795482 79 - 11822988 GUCUAUAAAAUGAUCUGCAAAUUAAAGCGUCUAAAGGCGCAGACGCCAGGCCGAC--CCAAAGA----------CAGAGCCCCCAACCGAC----------- (((..........(((((........))((((...((((....)))).((.....--))..)))----------))))..........)))----------- ( -14.55, z-score = -0.82, R) >droWil1.scaffold_181089 1807681 101 + 12369635 GUCUAUAAAAUUAUCUGCAAAUUAAAGCGUCUAA-CUCAGAAAAAAAAAAAUAAAAACCGCAAACGGCUAGUCUCAGAGCCAUUGAAAGACCAAAGGCAACC ((((............((........))((((..-.((((...................(....)((((........)))).)))).))))...)))).... ( -12.80, z-score = -0.65, R) >consensus GUCUAUAAAAUUAUCCGCAAAUUAAAGCGUCUAAAGCCAAAGACGCUAGCUGAAAAGCCGAAAA______AACGCAGAGCCACUGAAAAACCAAAAGACA__ ..........................((((((........))))))........................................................ ( -6.22 = -6.73 + 0.50)

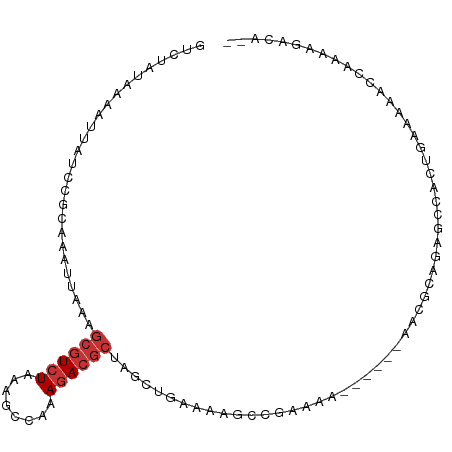

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:22:15 2011