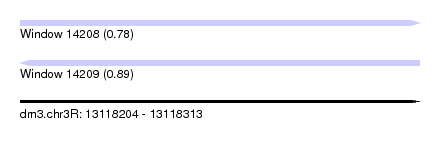

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,118,204 – 13,118,313 |
| Length | 109 |
| Max. P | 0.893338 |

| Location | 13,118,204 – 13,118,313 |
|---|---|
| Length | 109 |
| Sequences | 12 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 67.40 |
| Shannon entropy | 0.68611 |
| G+C content | 0.53588 |
| Mean single sequence MFE | -34.43 |
| Consensus MFE | -6.69 |
| Energy contribution | -6.70 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.19 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.780820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 13118204 109 + 27905053 ---GCUUCUGA---CUCCCUGGAUACUUUCCACGGAGUCAAGAGCAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCCCC-ACCAUCUGACAUAUUUGACUUCGCGUGCUCUUU ---(((..(((---((((.((((.....)))).)))))))..))).......((((((.....(((.((..((.((((....-.)))).))...)).))).....))))))..... ( -39.70, z-score = -4.71, R) >dp4.chr2 21484211 106 - 30794189 ---GCCACUG------CCGCGGAUACCUCAAGCGGAUUCAAGAGCAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCACC-ACCAUCUGACUUAUUGCCCUUCGUGUGCUCUUU ---(((..((------((..((((.((......))))))..).)))..)))((((((...(.(((((.((.((.((((....-.)))).)).)).))))))...))))))...... ( -30.80, z-score = -1.00, R) >droPer1.super_0 8705454 106 - 11822988 ---GCCACUG------CCGCGGAUACCUCAAGCGGAUUCAAGAGCAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCACC-ACCAUCUGACUUAUUGCCCUUCGUGUGCUCUUU ---(((..((------((..((((.((......))))))..).)))..)))((((((...(.(((((.((.((.((((....-.)))).)).)).))))))...))))))...... ( -30.80, z-score = -1.00, R) >droEre2.scaffold_4770 9237096 108 - 17746568 ---GCCGCUGA---CUCCCUGGUUGC-UUCCACGGACUCAAGAGCAACGGCCGCACGUCCGUGCAAUUAGCCAAAUGGCCCC-ACCAUCUGAGUUAUUUGACUUCGCGUGCUCGUG ---(((..(((---.(((.(((....-..))).))).))).).)).((((..((((((.....(((.((((((.((((....-.)))).)).)))).))).....)))))))))). ( -34.00, z-score = -1.37, R) >droYak2.chr3R 17208667 109 + 28832112 ---GCUCCUGA---CUCCCUGGACGCCUUCCACGGAGUCAAGAACAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCCCC-ACCAUCUGAGUUAUUUGACUUCGCAUGCUCUUU ---((((.(((---((((.((((.....)))).))))))).))......((.((((....))))...((((((.((((....-.)))).)).)))).........))..))..... ( -33.00, z-score = -2.52, R) >droSec1.super_12 1095471 109 - 2123299 ---GCUCCUGA---CUCCCUGGAUACUUUCCACGGAGUCAAGAGCAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCCCC-ACCAUCUGACAUAUUUGACUUCGGCUGCUCUUU ---((((.(((---((((.((((.....)))).))))))).))))..(((((((((....)))).....(((....)))...-......................)))))...... ( -43.70, z-score = -5.87, R) >droSim1.chr3R 19163904 109 - 27517382 ---GCUCUUGA---CUCCCUGGAUACUUUCCACGGAGUCAAGAGCAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCCCC-ACCAUCUGACAUAUUUGACUUUGGGUGCUCUUU ---((((((((---((((.((((.....)))).))))))))))))...(((.((((....)))).....)))(((.((((((-(...((..........))...)))).))).))) ( -46.20, z-score = -6.09, R) >droMoj3.scaffold_6540 17638564 99 + 34148556 CUAGGCGCUGGGUCAUGCCCCAACAUCGUCGGCGUGCC----GUCGAUGUCAACACGUUCGUGCAAUUAGCCAAAUGGCUCC-ACGGUCUGACUUGUGCCCUUU------------ ...(((((.((((((.(((...((((((.(((....))----).))))))..........(((.....((((....)))).)-))))).)))))))))))....------------ ( -39.00, z-score = -2.82, R) >droVir3.scaffold_13047 14214183 93 + 19223366 ---GCCGCAGGGUCAUGUCCUGGCAAC---GGCGUGCC----GUCGAUGCCUGCACGUACUUGCAAUUAGCCAAAUGGCUCC-ACGGGCUGACUUGUAUCCUUU------------ ---...(((((.(((.((((.((((.(---((((...)----)))).))))((((......))))...((((....))))..-..)))))))))))).......------------ ( -32.40, z-score = -0.52, R) >droGri2.scaffold_14830 530967 93 - 6267026 ---GCUGUUGGGUCAUGUCCUGACAUC---GGCGUGUC----GUCGAUGUCUGCACGUUCUUGUAAUUAGCCAAAUGGCUCC-ACGAUCUGACUUGUGUCUCUU------------ ---((.((..((((((((.(.((((((---((((...)----))))))))).).))))..........((((....))))..-..))))..))..)).......------------ ( -31.40, z-score = -2.97, R) >droWil1.scaffold_181089 1718051 103 + 12369635 ---GCAGCAGAA---UGCUUGUAUUCACCCUGUAGAUUCAAGAGCAAUGCCCGCACGUUCGUGCAAUUAGCCAAAUGGCCUCAACAGUCUGACUUUUUGCCUGCCCUUU------- ---(((((((((---((....)))))......((((((...(((........((((....)))).....(((....))))))...))))))......)).)))).....------- ( -25.10, z-score = -0.82, R) >droAna3.scaffold_13340 10334703 113 + 23697760 ---GCUCCCGGUUUCUCCCUGGAUAUUUUCGACGAAGUCAAGAGCAAUGGCCGCACGUUCUUGCAAUUAGCCAAAUGGCCCCCAACAUCUGACUUAUUGUUUUCCGUGUGUUUCCG ---((((((((.......))))........(((...)))..))))...((.((((((.....(((((.((.((.(((........))).)).)).)))))....))))))...)). ( -27.10, z-score = -1.10, R) >consensus ___GCUGCUGA___CUCCCUGGAUACCUUCCACGGAGUCAAGAGCAACGGCCGCACGUUCGUGCAAUUAGCCAAAUGGCCCC_ACCAUCUGACUUAUUUCACUUCGUGUGCUCUUU ....................................................(((......))).....(((....)))..................................... ( -6.69 = -6.70 + 0.01)



| Location | 13,118,204 – 13,118,313 |
|---|---|
| Length | 109 |
| Sequences | 12 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 67.40 |
| Shannon entropy | 0.68611 |
| G+C content | 0.53588 |
| Mean single sequence MFE | -39.59 |
| Consensus MFE | -8.37 |
| Energy contribution | -8.37 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.62 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.21 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.893338 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 13118204 109 - 27905053 AAAGAGCACGCGAAGUCAAAUAUGUCAGAUGGU-GGGGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGCUCUUGACUCCGUGGAAAGUAUCCAGGGAG---UCAGAAGC--- .(((((((.(((..(((......(((((((((.-....))))))))).....((((....)))))))))))))))))((((((.((((.....)))).))))---))......--- ( -49.50, z-score = -5.34, R) >dp4.chr2 21484211 106 + 30794189 AAAGAGCACACGAAGGGCAAUAAGUCAGAUGGU-GGUGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGCUCUUGAAUCCGCUUGAGGUAUCCGCGG------CAGUGGC--- .....((.(((.(((((((((.((((((((((.-....))))))))))..((((((....))))))..)))))))))....((((..((....)).))))------..)))))--- ( -40.90, z-score = -1.90, R) >droPer1.super_0 8705454 106 + 11822988 AAAGAGCACACGAAGGGCAAUAAGUCAGAUGGU-GGUGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGCUCUUGAAUCCGCUUGAGGUAUCCGCGG------CAGUGGC--- .....((.(((.(((((((((.((((((((((.-....))))))))))..((((((....))))))..)))))))))....((((..((....)).))))------..)))))--- ( -40.90, z-score = -1.90, R) >droEre2.scaffold_4770 9237096 108 + 17746568 CACGAGCACGCGAAGUCAAAUAACUCAGAUGGU-GGGGCCAUUUGGCUAAUUGCACGGACGUGCGGCCGUUGCUCUUGAGUCCGUGGAA-GCAACCAGGGAG---UCAGCGGC--- .(((.((.((((..(((........(((((((.-....)))))))((.....))...))).))))))))).(((.((((.(((.(((..-....))).))).---)))).)))--- ( -40.40, z-score = -1.36, R) >droYak2.chr3R 17208667 109 - 28832112 AAAGAGCAUGCGAAGUCAAAUAACUCAGAUGGU-GGGGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGUUCUUGACUCCGUGGAAGGCGUCCAGGGAG---UCAGGAGC--- ...(((.................))).((((((-..((((....))))...(((((....)))))))))))((((((((((((.((((.....)))).))))---))))))))--- ( -44.63, z-score = -3.32, R) >droSec1.super_12 1095471 109 + 2123299 AAAGAGCAGCCGAAGUCAAAUAUGUCAGAUGGU-GGGGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGCUCUUGACUCCGUGGAAAGUAUCCAGGGAG---UCAGGAGC--- ....(((.(((............(((((((((.-....))))))))).....((((....))))))).)))((((((((((((.((((.....)))).))))---))))))))--- ( -54.10, z-score = -6.42, R) >droSim1.chr3R 19163904 109 + 27517382 AAAGAGCACCCAAAGUCAAAUAUGUCAGAUGGU-GGGGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGCUCUUGACUCCGUGGAAAGUAUCCAGGGAG---UCAAGAGC--- ...(.((.((((..(((..........)))..)-))))))....((((....((((....))))))))...((((((((((((.((((.....)))).))))---))))))))--- ( -50.70, z-score = -5.84, R) >droMoj3.scaffold_6540 17638564 99 - 34148556 ------------AAAGGGCACAAGUCAGACCGU-GGAGCCAUUUGGCUAAUUGCACGAACGUGUUGACAUCGAC----GGCACGCCGACGAUGUUGGGGCAUGACCCAGCGCCUAG ------------...((((.(..((((...(((-(.((((....)))).....))))...((.(..((((((.(----((....))).))))))..).)).))))...).)))).. ( -39.30, z-score = -2.71, R) >droVir3.scaffold_13047 14214183 93 - 19223366 ------------AAAGGAUACAAGUCAGCCCGU-GGAGCCAUUUGGCUAAUUGCAAGUACGUGCAGGCAUCGAC----GGCACGCC---GUUGCCAGGACAUGACCCUGCGGC--- ------------..(((...((.(((.(((...-..((((....))))...((((......)))))))..((((----((....))---))))....))).))..))).....--- ( -27.80, z-score = 0.66, R) >droGri2.scaffold_14830 530967 93 + 6267026 ------------AAGAGACACAAGUCAGAUCGU-GGAGCCAUUUGGCUAAUUACAAGAACGUGCAGACAUCGAC----GACACGCC---GAUGUCAGGACAUGACCCAACAGC--- ------------....(((....)))......(-((((((....))))...........(((((.(((((((.(----(...)).)---))))))..).))))..))).....--- ( -24.10, z-score = -1.94, R) >droWil1.scaffold_181089 1718051 103 - 12369635 -------AAAGGGCAGGCAAAAAGUCAGACUGUUGAGGCCAUUUGGCUAAUUGCACGAACGUGCGGGCAUUGCUCUUGAAUCUACAGGGUGAAUACAAGCA---UUCUGCUGC--- -------.((((((((((....(((((((..((....))..)))))))..((((((....)))))))).)))))))).......(((((((........))---)))))....--- ( -31.20, z-score = -0.89, R) >droAna3.scaffold_13340 10334703 113 - 23697760 CGGAAACACACGGAAAACAAUAAGUCAGAUGUUGGGGGCCAUUUGGCUAAUUGCAAGAACGUGCGGCCAUUGCUCUUGACUUCGUCGAAAAUAUCCAGGGAGAAACCGGGAGC--- .((...(.((((.....((((.(((((((((........))))))))).))))......)))).).))...(((((((.((((.(.((.....)).).))))....)))))))--- ( -31.50, z-score = -0.96, R) >consensus AAAGAGCACACGAAGGGAAAUAAGUCAGAUGGU_GGGGCCAUUUGGCUAAUUGCACGAACGUGCGGCCGUUGCUCUUGACUCCGCGGAAGGUAUCCAGGGAG___UCAGCAGC___ ...........................((((.....((((....))))..(((((......))))).........................))))..................... ( -8.37 = -8.37 + -0.01)


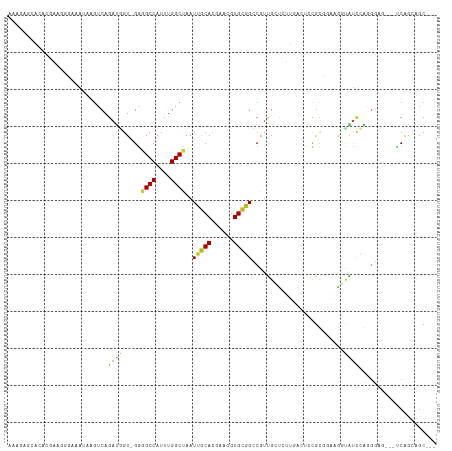
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:21:57 2011