
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,086,296 – 13,086,396 |
| Length | 100 |
| Max. P | 0.766230 |

| Location | 13,086,296 – 13,086,396 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 86.56 |
| Shannon entropy | 0.25704 |
| G+C content | 0.51667 |
| Mean single sequence MFE | -27.96 |
| Consensus MFE | -18.42 |
| Energy contribution | -19.15 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.766230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
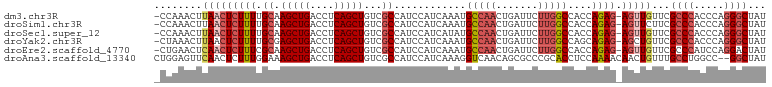
>dm3.chr3R 13086296 100 - 27905053 -CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUGUUCGCCCACCCAGGGCUAU -.......(((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-)))))...((((.....))))... ( -30.00, z-score = -3.19, R) >droSim1.chr3R 19131527 100 + 27517382 -CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUCUUCGCCCACCCAGGGCUAU -........((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-))))....((((.....))))... ( -28.50, z-score = -3.10, R) >droSec1.super_12 1052771 100 + 2123299 -CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAUAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUGUUCGCCCACCCAGGGCUAU -.......(((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-)))))...((((.....))))... ( -29.80, z-score = -3.17, R) >droYak2.chr3R 17172402 100 - 28832112 -CUAAACUUAACUCUUUUGCGAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCAGCAGAG-AGCUGUUCGCCCACCCAGGGCUAU -..........((((((.((.(((((....)))))...((((...((((.........))))...))))..))))))-))......((((.....))))... ( -29.50, z-score = -1.95, R) >droEre2.scaffold_4770 9204948 100 + 17746568 -CUGAACUCAACUCUUUCGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUGUUCGCCCAUCCAGGACUAU -..((((..((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-))))))))..((.....))..... ( -25.80, z-score = -1.80, R) >droAna3.scaffold_13340 10303792 100 - 23697760 CUGGAGUUCAACUCUUUGGAAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAGGUCAACAGCGCCCGCACCUCCAAAACAACUGUUUGCCUGGCC--GGCUAU ..((((.....))))...((.(((((....))))).))(((..(((.....(((.(((((....................))))).))))))..--)))... ( -24.15, z-score = 0.26, R) >consensus _CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG_AGUUGUUCGCCCACCCAGGGCUAU ....((((...((((...((.(((((....)))))...))............(((((.......)))))....)))).))))....((((.....))))... (-18.42 = -19.15 + 0.73)


| Location | 13,086,297 – 13,086,396 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 86.42 |
| Shannon entropy | 0.25958 |
| G+C content | 0.52189 |
| Mean single sequence MFE | -27.96 |
| Consensus MFE | -18.42 |
| Energy contribution | -19.15 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.665948 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
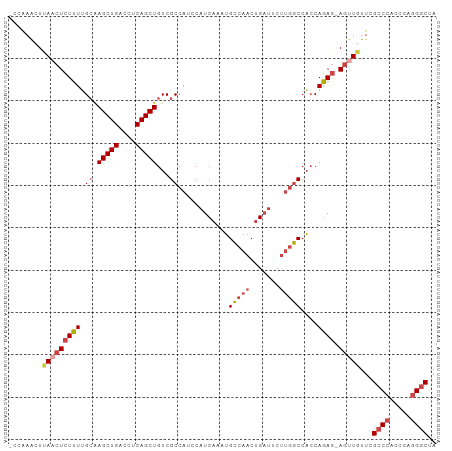
>dm3.chr3R 13086297 99 - 27905053 -CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUGUUCGCCCACCCAGGGCUA -.......(((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-)))))...((((.....)))).. ( -30.00, z-score = -2.97, R) >droSim1.chr3R 19131528 99 + 27517382 -CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUCUUCGCCCACCCAGGGCUA -........((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-))))....((((.....)))).. ( -28.50, z-score = -2.94, R) >droSec1.super_12 1052772 99 + 2123299 -CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAUAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUGUUCGCCCACCCAGGGCUA -.......(((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-)))))...((((.....)))).. ( -29.80, z-score = -3.00, R) >droYak2.chr3R 17172403 99 - 28832112 -CUAAACUUAACUCUUUUGCGAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCAGCAGAG-AGCUGUUCGCCCACCCAGGGCUA -..........((((((.((.(((((....)))))...((((...((((.........))))...))))..))))))-))......((((.....)))).. ( -29.50, z-score = -1.83, R) >droEre2.scaffold_4770 9204949 99 + 17746568 -CUGAACUCAACUCUUUCGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG-AGUUGUUCGCCCAUCCAGGACUA -..((((..((((((((....(((((....)))))((.((((...((((.........))))...)))).)).))))-))))))))..((.....)).... ( -25.80, z-score = -1.76, R) >droAna3.scaffold_13340 10303793 99 - 23697760 CUGGAGUUCAACUCUUUGGAAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAGGUCAACAGCGCCCGCACCUCCAAAACAACUGUUUGCCUGGCC--GGCUA ..((((.....))))...((.(((((....))))).))(((..(((.....(((.(((((....................))))).))))))..--))).. ( -24.15, z-score = 0.39, R) >consensus _CCAAACUUAACUCUUUUGCAAGCUGACCUCAGCUGUCGCCAUCCAUCAAAUGCCAACUGAUUCUUGGCCACCAGAG_AGUUGUUCGCCCACCCAGGGCUA ....((((...((((...((.(((((....)))))...))............(((((.......)))))....)))).))))....((((.....)))).. (-18.42 = -19.15 + 0.73)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:21:50 2011