
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 13,072,015 – 13,072,129 |
| Length | 114 |
| Max. P | 0.545191 |

| Location | 13,072,015 – 13,072,129 |
|---|---|
| Length | 114 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 74.30 |
| Shannon entropy | 0.49161 |
| G+C content | 0.41248 |
| Mean single sequence MFE | -28.21 |
| Consensus MFE | -10.88 |
| Energy contribution | -10.73 |
| Covariance contribution | -0.15 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.545191 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 13072015 114 - 27905053 CUUGCCCAUCCUCCGUGGUC--CUCCUGUAUGGCUUGAUUAUUUCGUUU-GGAAUUGAUGCUGCCAUCAAUUAUAGAAAACCACAAAGAGAAGCAAGACAGCAGGAUGGGAAAAAUG ....((((((((..(((((.--...(((((((((...((((.(((....-.))).))))...)))).....)))))...)))))........((......))))))))))....... ( -33.10, z-score = -2.03, R) >droSim1.chr3R 19115097 114 + 27517382 CUUGCCCAUCCUCCGUGGUC--CUCUUGUAUGGCUUGAUUAUUUCGUUU-GGAAUUGAUGCUGCCAUCAAUUAUAGAAAACCACAAAGAGGAGCAAGACAGCAGGAUGGGAAAAAUG ....((((((((.(.((.((--(((((...(((...((.....)).(((-(.((((((((....)))))))).))))...)))..))))))).)).....).))))))))....... ( -37.70, z-score = -3.00, R) >droSec1.super_12 1038829 114 + 2123299 CUUGCUCAUCCUCCGUGGUC--CUCUUGUAUGGCUUGAUUAUUUCGUUU-GGAAUUGAUGCUGCCAUCAAUUAUAGAAAACCACAAAGAGGAGCAAGACAGCAGGAUGGGAAAAAUG ....((((((((.(.((.((--(((((...(((...((.....)).(((-(.((((((((....)))))))).))))...)))..))))))).)).....).))))))))....... ( -34.70, z-score = -2.18, R) >droYak2.chr3R 17157726 102 - 28832112 ------------CCAUGGUC--CUCCUGUAUGGCUUGAUUAUUUCGUUU-GGAAUUGAUGCUGCCAUCAAUUAUAGAAAACCACAAAGAGGAGCAAGACAGCAGGAUGGGAAAAAUG ------------((...(((--((.((((...((((((.....))((((-..((((((((....)))))))).....)))).........))))...)))).))))).))....... ( -26.40, z-score = -1.06, R) >droEre2.scaffold_4770 9190744 102 + 17746568 ------------CCAUGGUC--CUCCUGCAUGGCUUGAUUAUUUCGUUU-GGAAUUGAUGCUGCCAUCAAUUAUAGAAAACCACAAAGAGGAGCAGGACAGCAGGAUGGGAAAAAUG ------------.(((..((--(((((((.((.((((....((((....-..((((((((....))))))))...)))).((.......))..)))).))))))))..)))...))) ( -27.80, z-score = -0.98, R) >droAna3.scaffold_13340 10290712 108 - 23697760 --------UCUCCCAUGACCAAAUUCUGUAUUGCUUGAUUAUUUCGUUU-GAAAUUGAUGCUGCCAUCAAUUACAGAAAACCACAAAGACGACCCAGACAGUGGAAUGGGAAAAAUG --------..((((((..(((...((((..((((((..........(((-(.((((((((....)))))))).))))........))).)))..))))...))).))))))...... ( -30.47, z-score = -4.00, R) >droGri2.scaffold_14830 485079 88 + 6267026 -----------------------GUUAGAGUUGUUUGAUUAUUUCGUUUGGAAAUGGAUGCUGCAAUCAAUUAUAGAGCACCACAAAG----CGAAGAGAGAAGACAGA--GAAUCG -----------------------....((.((((((..((.((((((((.....(((.((((..............)))))))..)))----))))).))..)))))).--...)). ( -19.24, z-score = -1.62, R) >droMoj3.scaffold_6540 17587466 93 - 34148556 -----------------------GUGUGCACUGCUUGAUUAUUUCGUUUUGAAAUUGAUGCUGCAAUCAAUUAUAGAAGAACGCAAAAA-ACCAGAGAGAACAAGUAGAAUGAAUCG -----------------------.......(((((((....((((.(((((.(((((((......))))))).)))))(....).....-....))))...)))))))......... ( -16.30, z-score = -0.15, R) >consensus ____________CCAUGGUC__CUCCUGUAUGGCUUGAUUAUUUCGUUU_GGAAUUGAUGCUGCCAUCAAUUAUAGAAAACCACAAAGAGGAGCAAGACAGCAGGAUGGGAAAAAUG .......................((((((....((((....(((.((((...((((((((....)))))))).....))))...)))......))))...))))))........... (-10.88 = -10.73 + -0.15)
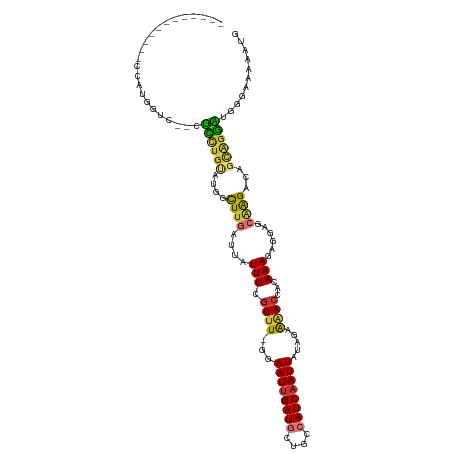


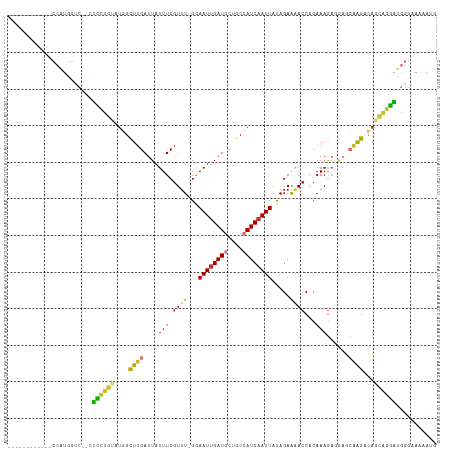
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:21:48 2011