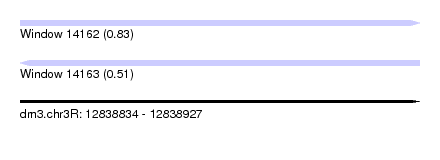
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 12,838,834 – 12,838,927 |
| Length | 93 |
| Max. P | 0.833294 |

| Location | 12,838,834 – 12,838,927 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 80.47 |
| Shannon entropy | 0.37061 |
| G+C content | 0.46040 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -14.97 |
| Energy contribution | -16.80 |
| Covariance contribution | 1.83 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.833294 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
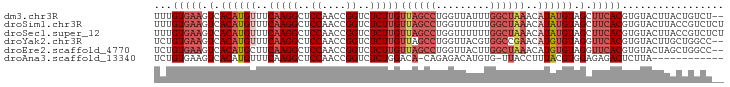
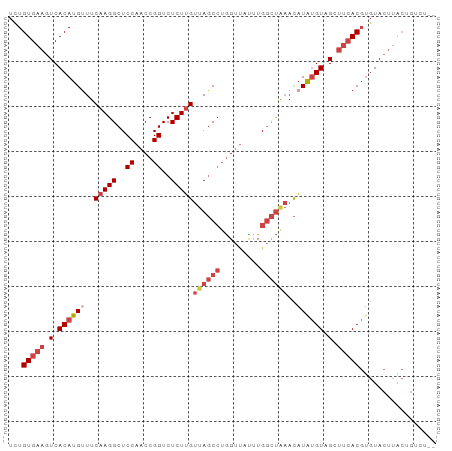
>dm3.chr3R 12838834 93 + 27905053 UUUGUGAAGUCACAUGUUUCAAGGCUCCAACCGGUCUCUUGUUAGCCUGGUUAUUUGGCUAAACAUAUGUAGCUUCACGUGUACUUACUGUCU-- ...((((((..((((((..(((((..((....))..)))))((((((.........))))))..))))))..))))))(((....))).....-- ( -28.50, z-score = -3.37, R) >droSim1.chr3R 18882974 95 - 27517382 UUUGUGAAGUCACAUGUUUCAAGGCUCCAACCGGUCUCUUGUUAGCCUGGUUUUUUGGCUAAACAUAUGUAGCUUCACGUGUACUUACCGUCUCU ...((((((..((((((..(((((..((....))..)))))((((((.........))))))..))))))..))))))................. ( -28.30, z-score = -3.40, R) >droSec1.super_12 806006 95 - 2123299 UUUGUGAAGUCACAUGUUUCAAGGCUCCAACCGGUCUCUUGUUAGCCUGGUUUUUUGGCUAAACAUAUGUAGCUUCACGUGUACUUACCGUCUCU ...((((((..((((((..(((((..((....))..)))))((((((.........))))))..))))))..))))))................. ( -28.30, z-score = -3.40, R) >droYak2.chr3R 17286926 93 - 28832112 UCUGUGAAGUCACAUGUUUCAAGGCUCCAACCGGUCUCUUGUUAGCCUGGUUACGUGGCCGAACAUGUGUAGGUUCACGUGUACUUGCUGGCC-- ...(((((.((((((((((...(((..(((((((.((......)).))))))..)..))))))))))))..).)))))...............-- ( -27.90, z-score = -0.65, R) >droEre2.scaffold_4770 8954688 93 - 17746568 UCUGUGAAGUCACAUGCUUCAAGGCUCCAACCGGUCUCUUGUUAGCCUGGUUACUUGGCUAAACAUGUGUAGGUUCACGUGUACUAGCUGGCC-- ....((((((.....)))))).((((..((((((.((......)).))))))....(((((.(((((((......)))))))..)))))))))-- ( -28.90, z-score = -1.34, R) >droAna3.scaffold_13340 16633671 81 - 23697760 UCUGUGAAGUCACAUGUUUCAAGGCUCCAACCGGUCUCUGGACA-CAGAGACAUGUG-UUACCUUUACGUGGAGAGACUCUUA------------ ...((((((.((((((((((..((......)).(((....))).-..))))))))))-....))))))..((((...))))..------------ ( -24.10, z-score = -0.88, R) >consensus UCUGUGAAGUCACAUGUUUCAAGGCUCCAACCGGUCUCUUGUUAGCCUGGUUAUUUGGCUAAACAUAUGUAGCUUCACGUGUACUUACUGUCU__ ...(((((.(.((((((..(((((..((....))..)))))((((((.........))))))..)))))).).)))))................. (-14.97 = -16.80 + 1.83)

| Location | 12,838,834 – 12,838,927 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 80.47 |
| Shannon entropy | 0.37061 |
| G+C content | 0.46040 |
| Mean single sequence MFE | -23.99 |
| Consensus MFE | -15.32 |
| Energy contribution | -16.72 |
| Covariance contribution | 1.39 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.510055 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 12838834 93 - 27905053 --AGACAGUAAGUACACGUGAAGCUACAUAUGUUUAGCCAAAUAACCAGGCUAACAAGAGACCGGUUGGAGCCUUGAAACAUGUGACUUCACAAA --...............((((((...(((((((((.((....(((((.((((......)).)))))))..))....))))))))).))))))... ( -24.80, z-score = -2.61, R) >droSim1.chr3R 18882974 95 + 27517382 AGAGACGGUAAGUACACGUGAAGCUACAUAUGUUUAGCCAAAAAACCAGGCUAACAAGAGACCGGUUGGAGCCUUGAAACAUGUGACUUCACAAA .................((((((...(((((((((((((.........))))..((((...((....))...))))))))))))).))))))... ( -24.50, z-score = -2.05, R) >droSec1.super_12 806006 95 + 2123299 AGAGACGGUAAGUACACGUGAAGCUACAUAUGUUUAGCCAAAAAACCAGGCUAACAAGAGACCGGUUGGAGCCUUGAAACAUGUGACUUCACAAA .................((((((...(((((((((((((.........))))..((((...((....))...))))))))))))).))))))... ( -24.50, z-score = -2.05, R) >droYak2.chr3R 17286926 93 + 28832112 --GGCCAGCAAGUACACGUGAACCUACACAUGUUCGGCCACGUAACCAGGCUAACAAGAGACCGGUUGGAGCCUUGAAACAUGUGACUUCACAGA --...............(((((....((((((((.(((..((..(((.((((......)).)))))))..)))....))))))))..)))))... ( -23.90, z-score = -0.37, R) >droEre2.scaffold_4770 8954688 93 + 17746568 --GGCCAGCUAGUACACGUGAACCUACACAUGUUUAGCCAAGUAACCAGGCUAACAAGAGACCGGUUGGAGCCUUGAAGCAUGUGACUUCACAGA --((((.(((((.(((.(((......))).))))))))....(((((.((((......)).)))))))).))).((((((....).))))).... ( -23.80, z-score = -0.54, R) >droAna3.scaffold_13340 16633671 81 + 23697760 ------------UAAGAGUCUCUCCACGUAAAGGUAA-CACAUGUCUCUG-UGUCCAGAGACCGGUUGGAGCCUUGAAACAUGUGACUUCACAGA ------------...(((((.(........(((((..-(((..(((((((-....)))))))..).))..))))).......).)))))...... ( -22.46, z-score = -0.74, R) >consensus __AGACAGUAAGUACACGUGAAGCUACAUAUGUUUAGCCAAAUAACCAGGCUAACAAGAGACCGGUUGGAGCCUUGAAACAUGUGACUUCACAAA .................(((((....(((((((((((((.........))))..((((...((....))...)))))))))))))..)))))... (-15.32 = -16.72 + 1.39)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:21:17 2011