
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 12,695,121 – 12,695,213 |
| Length | 92 |
| Max. P | 0.547486 |

| Location | 12,695,121 – 12,695,213 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 59.30 |
| Shannon entropy | 0.82672 |
| G+C content | 0.48760 |
| Mean single sequence MFE | -25.92 |
| Consensus MFE | -5.30 |
| Energy contribution | -4.69 |
| Covariance contribution | -0.61 |
| Combinations/Pair | 1.87 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.20 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.547486 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
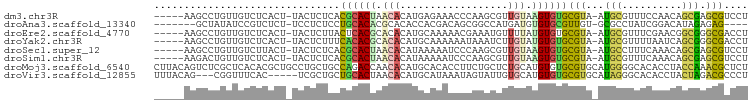
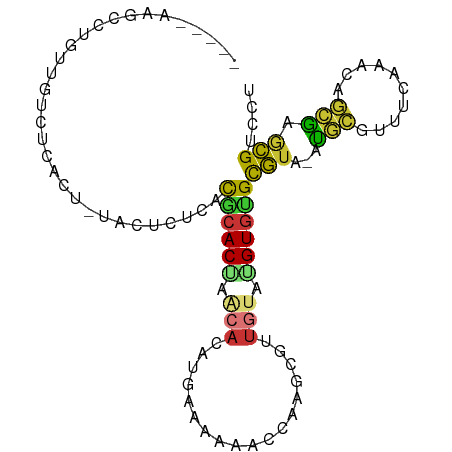
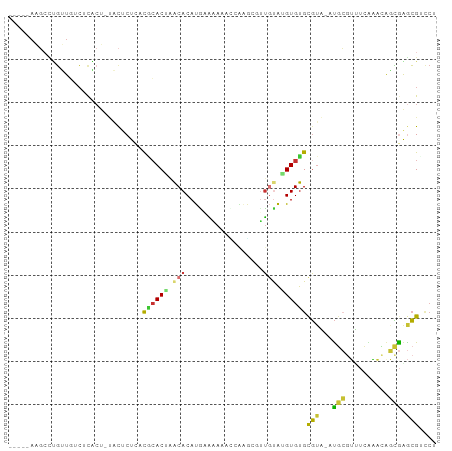
>dm3.chr3R 12695121 92 - 27905053 -----AAGCCUGUUGUCUCACU-UACUCUCACGCACUAACACAUGAGAAACCCAAGCGUUGUAAGUGUGCGUA-AUGCGUUUCCAACAGCGAGCGUCCU -----..((.((((((..((((-((((((((.(........).)))))(((......)))))))))).(((..-...))).....)))))).))..... ( -20.60, z-score = -0.22, R) >droAna3.scaffold_13340 16495167 86 + 23697760 -------GCUAUAUCCGUCUCU-UCCUCUCCUGCACACGCACACCACGACAGCGGCCAUGAUGUGUGCGUUGU-GCGCCUAUCGGACAUAGAGAG---- -------..........(((((-(((.....(((((((((((((((.(........).))..))))))).)))-)))......)))...))))).---- ( -25.30, z-score = -0.80, R) >droEre2.scaffold_4770 8807686 92 + 17746568 -----AAGCCUGUUGUCUCACU-UACUCUUACUCACGCACACAUGCAAAAACGAAAUGUUUUAUGUGUGCGUA-AUGCGUUUCGAACGGCGGGCGACCU -----..(((((((((.((...-.((.(......(((((((((((...((((.....))))))))))))))).-..).))...)))))))))))..... ( -29.90, z-score = -2.94, R) >droYak2.chr3R 17136562 92 + 28832112 -----AAGCCUGUUGUCUCACU-UACUCUUUCACACGCACACAUGCAAAAAAUAAAUCUUGUAUGUGUGCGUA-AUGCGUUUUAAUCAGCGGGCGACCU -----..((((((((.......-.((.(......((((((((((((((..........)))))))))))))).-..).))......))))))))..... ( -32.74, z-score = -5.27, R) >droSec1.super_12 664008 92 + 2123299 -----AAGCCUGUUGUCUUACU-UACUCUCACGCACUAACACAUAAAAAUCCCAAGCGUUGUAAGUGUGCGUA-AUGCCUUUCAAACAGCGAGCGUCCU -----..((.((((((..((((-(((....((((.....................)))).))))))).((...-..)).......)))))).))..... ( -16.90, z-score = -0.51, R) >droSim1.chr3R 18739549 92 + 27517382 -----AAGACUGUUGUCUCACU-UACUCUCACGCACUAACACAUAAAAAUCCCAAGCGUUGUAAGUGUGCGUA-AUGCGUUUCAAACAGCGAGCGUCCU -----.((((....))))((((-(((....((((.....................)))).))))))).((((.-...((((......)))).))))... ( -17.90, z-score = -0.20, R) >droMoj3.scaffold_6540 1960563 99 - 34148556 CUUACAGUCUCGCUCACACGCUGCCUGCUGCCAGACCAACACAUGCACACCUUCUGCUCUGCAUGUGUGCGUGCAUGGGGCACACCUACCAAACGCUCU ......((((.((......((.....)).)).))))..(((((((((...(....)...)))))))))((((...((((.....))))....))))... ( -26.70, z-score = -0.24, R) >droVir3.scaffold_12855 9763657 91 + 10161210 UUUACAG---CGGUUUCAC-----UCGCUGCUGCACUAACACAUGCAUAAAUAGUAUUGUGCAUGUGUGCGUGCAUAGGGCACACCUACUAGACGCCCU ....(((---(((......-----)))))).(((((..((((((((((((......))))))))))))..))))).(((((.............))))) ( -37.32, z-score = -3.94, R) >consensus _____AAGCCUGUUGUCUCACU_UACUCUCACGCACUAACACAUGAAAAAACCAAGCGUUGUAUGUGUGCGUA_AUGCGUUUCAAACAGCGAGCGUCCU ...............................((((((.(((..................))).))))))(((...(((..........))).))).... ( -5.30 = -4.69 + -0.61)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:20:59 2011