
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 12,576,538 – 12,576,637 |
| Length | 99 |
| Max. P | 0.682689 |

| Location | 12,576,538 – 12,576,637 |
|---|---|
| Length | 99 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 60.85 |
| Shannon entropy | 0.75531 |
| G+C content | 0.32799 |
| Mean single sequence MFE | -20.53 |
| Consensus MFE | -5.42 |
| Energy contribution | -5.86 |
| Covariance contribution | 0.43 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.682689 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 12576538 99 - 27905053 GGAAGAAAUACAAAGUUAUUGCUGCAAAAC---CUAAAAUUAAACUAGAAAUUUUAGAAACAAUUUCCU-ACAA------CUGGGUCAAGUUUUUAGGAAAUUGGUUCC ((((..........................---((((((((........))))))))...(((((((((-(.((------((......))))..)))))))))).)))) ( -21.20, z-score = -2.09, R) >droAna3.scaffold_13340 16386842 107 + 23697760 AGCGUAUUUAUGAAGUUUUUGUCGCGGAGCGA-CAGAGAUU-GCCUAAAUCUUUUAGGAAGAAUUUCCUGAUAGGUUGCCGCUUCGUAAUAUUUUGGGAUGUUAUUUCC .......((((((((((((((((((...))))-))))))))-((...(((((.(((((((....))))))).)))))...)).))))))......((((......)))) ( -31.10, z-score = -2.47, R) >droEre2.scaffold_4770 8692793 98 + 17746568 GCAAGAAAUGCAAAGCUAUUGCUGCAAAAC---UUAAAAUUAAACUAAAACUUUUGGGAACAAUUUCCU-ACAA------GUGGGUC-AAGUUUUAGGAAAUUGUUUCC ..(((...((((..((....))))))...)---))....................((((((((((((((-(.((------.(.....-.).)).))))))))))))))) ( -24.70, z-score = -2.71, R) >droYak2.chr3R 17020631 82 + 28832112 -------------------GGAGGAAAUACAAUUGGAAAUUAAACUACACCUUUUAGAAACAAUUUCCUAGAAG--------UGGGUCAAGUUUUAGGAAAUUGUUUCC -------------------(((((.........................)))))..(((((((((((((((((.--------(......).))))))))))))))))). ( -22.01, z-score = -2.74, R) >droSec1.super_12 552084 99 + 2123299 GGAAGAAACACAAAGUUAUUGCUGCAAAAC---CUAAAAUUAAACUAGACAUUUUAGGAGCGAUUUCCU-ACAA------CUGGGUUAAGUUUUUAGAAAAUUGGUUCC ((((.........................(---(((((((..........))))))))..((((((.((-(.((------((......))))..))).)))))).)))) ( -15.10, z-score = 0.43, R) >droSim1.chr3R 18623743 99 + 27517382 GGAAGAAACACAAAGUUAUUGCUGCAAAAC---CUAAAAUUAAACUAGACAUUUUAGGAGCGAUUUCCU-ACAA------CUGGGUCAAGUUUUUAGGAAAUUGGUUCC ((((.........................(---(((((((..........))))))))..(((((((((-(.((------((......))))..)))))))))).)))) ( -21.90, z-score = -1.53, R) >droWil1.scaffold_180701 2676849 97 + 3904529 GUAAUAAAUACAUAAUCUUUUUCGUAAAAC---CUAAAAAUCAUCUAAAAUAUUUAUGUAGUAAACUUU-AUAU------CCUUCUC-CGAUACCGGAUCAUCAUCUU- ........(((((((...((((.(......---...........).))))...))))))).........-....------.....((-((....))))..........- ( -7.73, z-score = -1.00, R) >consensus GGAAGAAAUACAAAGUUAUUGCUGCAAAAC___CUAAAAUUAAACUAAAAAUUUUAGGAACAAUUUCCU_ACAA______CUGGGUCAAGUUUUUAGGAAAUUGUUUCC ........................................................(((.(((((((((..........................))))))))).))). ( -5.42 = -5.86 + 0.43)


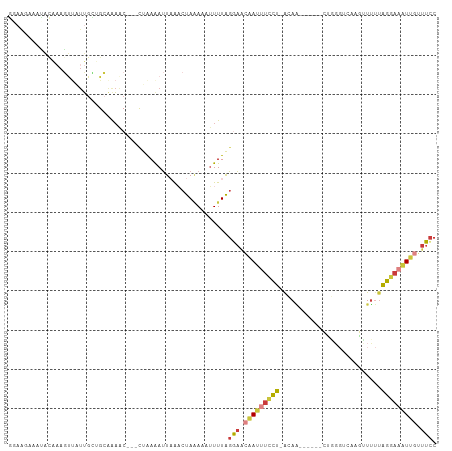
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:20:40 2011