
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 12,536,267 – 12,536,389 |
| Length | 122 |
| Max. P | 0.977261 |

| Location | 12,536,267 – 12,536,364 |
|---|---|
| Length | 97 |
| Sequences | 13 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 76.68 |
| Shannon entropy | 0.48562 |
| G+C content | 0.39876 |
| Mean single sequence MFE | -21.96 |
| Consensus MFE | -16.13 |
| Energy contribution | -15.95 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.13 |
| Mean z-score | -0.68 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 12536267 97 - 27905053 ------UUAUAAAC--GUUUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUCGAAGUUUUG--------- ------........--..((((.((.(((((...(((((((((((((.......)))))))))((...........))...))))..)))))))..)))).....--------- ( -19.80, z-score = -0.06, R) >droSim1.chr3R 18583217 97 + 27517382 ------UUAUAAAC--UUUUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAUAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUCGAAGUUUUG--------- ------....((((--(((..((...(((((...(((((((((((((.......)))))))))((...........))...))))..)))))))..)))))))..--------- ( -23.00, z-score = -1.93, R) >droSec1.super_12 512110 97 + 2123299 ------UUAUAAAC--UUUUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUCGAAGUUUUG--------- ------....((((--(((..((...(((((...(((((((((((((.......)))))))))((...........))...))))..)))))))..)))))))..--------- ( -23.00, z-score = -1.28, R) >droYak2.chr3R 16978533 97 + 28832112 ------UUAUAAAC--CAUUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUGAAAGUUUUG--------- ------........--..((((.((.(((((...(((((((((((((.......)))))))))((...........))...))))..))))))).))))......--------- ( -22.40, z-score = -0.98, R) >droEre2.scaffold_4770 8651901 97 + 17746568 ------UUAUAAAC--CUUUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUCAAAGUUUUG--------- ------...((((.--((((...((.(((((...(((((((((((((.......)))))))))((...........))...))))..)))))))..)))).))))--------- ( -20.40, z-score = -0.60, R) >droAna3.scaffold_13340 16348618 102 + 23697760 UCGCAACUAAACA---UCAUCUUCUUUCUGAAUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUCGAUGUUCUU--------- .........((((---((..(((...((((....(((((((((((((.......)))))))))((...........))...)))).)))).)))..))))))...--------- ( -25.40, z-score = -1.32, R) >dp4.chr2 17520006 104 - 30794189 -GCCGAUUAAACCCCGUAUUUCUCUUUCUGUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCUCUUCUACCAGACUGGCUAGGUAAGUCGACUUUUUG--------- -..(((((....(((............(((.....)))(((((((((.......))))))))))))........((((((.....)).)))))))))........--------- ( -23.90, z-score = -0.40, R) >droPer1.super_0 11369121 104 - 11822988 -GCCGAUUAAACGCAGUAUUUCUCUUUCUCUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCUCUUCUACCAGACUGGCUAGGUAAGUCGACUUUUUG--------- -..(((((...(((((...((((...((((((........)))))).....))))....)))))((((.(((......)))..)))).....)))))........--------- ( -21.20, z-score = 0.54, R) >droWil1.scaffold_181130 11580808 107 + 16660200 ----AUCACAUU---UUGUUAUUUCUAUUACUUUUCAGGUAAGAGAUACUCCGAAUCUCUUGCGGGCUCUCUUCUACCAGACUGGCUAGGUAAGUGCAACUUGUUAUUUCUUUU ----...(((..---((((........((((((.(((((((((((((.......)))))))))((...........))...))))..))))))..))))..))).......... ( -22.30, z-score = -1.21, R) >droVir3.scaffold_13047 13500192 104 + 19223366 UUUCAACUAAGC-AUAUGUUUUCUCUAUUAAUUUUCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGAUCACUUCUUCCAGACUGGCUAGGUAAGUGCACUUUUCU--------- .....(((.(((-....((((.................(((((((((.......))))))))).(((........)))))))..))).)))..............--------- ( -20.50, z-score = -0.22, R) >droMoj3.scaffold_6540 18025865 103 + 34148556 UCCCAACUAACC-AAAUGAUUU-UCUAUUUCUUUUCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCUCUUCUACCAGACUGGCUAGGUAAGUGCAAUGUUAU--------- ....(((...((-...(((...-...........))).(((((((((.......)))))))))))((.((..((((((.....)).))))..)).))...)))..--------- ( -21.94, z-score = -0.44, R) >droGri2.scaffold_14906 3087029 100 + 14172833 -----UCUCAUUUAAUUGGUUUCUCUUUUGUUUUUCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGAUCACUUCUACCAGACUGGCUAGGUAAGUGCAUUUUCCU--------- -----...((((((.((((((..(((............(((((((((.......)))))))))((...........)))))..)))))).)))))).........--------- ( -21.90, z-score = -0.40, R) >anoGam1.chr2R 61944072 91 - 62725911 ------CUAAAGA----AAUCUUCUUUUU----UAUAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCACUCUUCUACCAGACUGGAUAGGUAAGUU-ACGUUUUUA-------- ------(((.(((----((......))))----).)))(((((((((.......))))))))).(..(((..((((((.....)).))))..))).-.).......-------- ( -19.70, z-score = -0.59, R) >consensus ______UUAAAAAC__UAUUUCUCUUUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUCUACCAGACUGGCUAGGUAAGUCGAAGUUUUG_________ ..................................(((((((((((((.......)))))))))((...........))...))))............................. (-16.13 = -15.95 + -0.18)


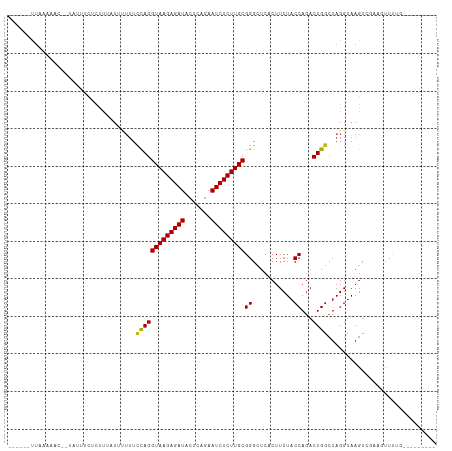
| Location | 12,536,298 – 12,536,389 |
|---|---|
| Length | 91 |
| Sequences | 13 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 65.61 |
| Shannon entropy | 0.72605 |
| G+C content | 0.39955 |
| Mean single sequence MFE | -19.43 |
| Consensus MFE | -11.29 |
| Energy contribution | -11.11 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.18 |
| Mean z-score | -0.92 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.97 |
| SVM RNA-class probability | 0.977261 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 12536298 91 - 27905053 --------------GAGGCAUAAUGACGUUCCUGGAC---CUUUAUAAACGU----UUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUC --------------(((.(............(((((.---............----...............)))))(((((((((.......))))))))).).)))..... ( -20.29, z-score = -0.95, R) >droSim1.chr3R 18583248 91 + 27517382 --------------GAGGCAUUAUUGCGUUCCUGGAC---CUUUAUAAACUU----UUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAUAAUCUCUUGCGGGCUCACUUC --------------(((.(............(((((.---............----...............)))))(((((((((.......))))))))).).)))..... ( -20.29, z-score = -2.08, R) >droSec1.super_12 512141 91 + 2123299 --------------GAGGCAAUAUUGCGUUCCUGGAC---CUUUAUAAACUU----UUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUC --------------(((.(............(((((.---............----...............)))))(((((((((.......))))))))).).)))..... ( -20.29, z-score = -1.29, R) >droYak2.chr3R 16978564 91 + 28832112 --------------GAGGCAUAAUCACGUUCCUGGAC---CUUUAUAAACCA----UUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUC --------------(((.(............(((((.---............----...............)))))(((((((((.......))))))))).).)))..... ( -20.29, z-score = -1.64, R) >droEre2.scaffold_4770 8651932 91 + 17746568 --------------GAGGCAUAAUGACGUUCCUUCGC---CUUUAUAAACCU----UUUCUCUCUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUC --------------(((.(((((.(.((......)).---).))))......----....................(((((((((.......))))))))).).)))..... ( -17.00, z-score = -0.50, R) >droAna3.scaffold_13340 16348649 94 + 23697760 --------------GAGGCAUU-UGUAGUCCUUGACCUUCGCAACUAAACAUC---AUCUUCUUUCUGAAUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUC --------------((((....-((.(((((......................---.........(((.....)))(((((((((.......)))))))))))))))))))) ( -20.30, z-score = -0.59, R) >dp4.chr2 17520037 98 - 30794189 --------------GAGGCAUUUUCCCGCCCAUCGACGUGCCGAUUAAACCCCGUAUUUCUCUUUCUGUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCUCUUC --------------((((.........((((...((.((((.(........).)))).)).....(((.....)))(((((((((.......)))))))))))))))))... ( -23.30, z-score = -1.03, R) >droPer1.super_0 11369152 98 - 11822988 --------------GAGGCAUUUUCCCGCCCAUCGACGUGCCGAUUAAACGCAGUAUUUCUCUUUCUCUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCUCUUC --------------((((.........((((...((.((((((......))..)))).))................(((((((((.......)))))))))))))))))... ( -22.50, z-score = -0.59, R) >droWil1.scaffold_181130 11580848 91 + 16660200 --------------AAAGUA-UUAUACAAUUUUGAAUUAAUCACAUU------UUGUUAUUUCUAUUACUUUUCAGGUAAGAGAUACUCCGAAUCUCUUGCGGGCUCUCUUC --------------((((((-....((((...(((.....)))....------)))).........))))))....(((((((((.......)))))))))........... ( -13.52, z-score = -0.78, R) >droVir3.scaffold_13047 13500223 111 + 19223366 AUAAUCAAAACGUUGAGGCA-UUUCUUAAUCUUAAACACUUUCAACUAAGCAUAUGUUUUCUCUAUUAAUUUUCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGAUCACUUC ............(((((..(-(((((((.(((.(((...........((((....))))...........))).))))))))))).))))).(((((....)))))...... ( -16.25, z-score = -0.46, R) >droMoj3.scaffold_6540 18025896 108 + 34148556 -AAAGCGAACCGUU-AGGCA-UUUGGCCAUAUUAAACUCUCCCAACUAACCAAAUGAUUU-UCUAUUUCUUUUCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCUCUUC -.(((.((.((...-...((-(((((.......................)))))))....-...............(((((((((.......))))))))).)).)).))). ( -20.80, z-score = -0.56, R) >droGri2.scaffold_14906 3087060 103 + 14172833 -----AGAAACUUUGAGGCAUUUAUUUAAUUUUGGA-CACUCUCAUU---UAAUUGGUUUCUCUUUUGUUUUUCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGAUCACUUC -----(((((((.(((((..((((........))))-..).))))..---.....))))))).....((..(((..(((((((((.......))))))))).)))..))... ( -20.50, z-score = -0.79, R) >anoGam1.chr2R 61944103 77 - 62725911 ---------------------------AUUGUGGAUCGAGAUAUCUAAAGAA----AUCUUCUUU----UUUAUAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCACUCUUC ---------------------------.(((((((..(((((.((....)).----)))))....----)))))))(((((((((.......)))))))))........... ( -17.20, z-score = -0.73, R) >consensus ______________GAGGCAUUUUCACGUUCUUGGAC___CUUUAUAAACCA____UUUCUCUUUAUUUUUUCCAGGUAAGAGAUACUCAGAAUCUCUUGCGGGCUCACUUC ........................................................................((..(((((((((.......))))))))).))........ (-11.29 = -11.11 + -0.18)

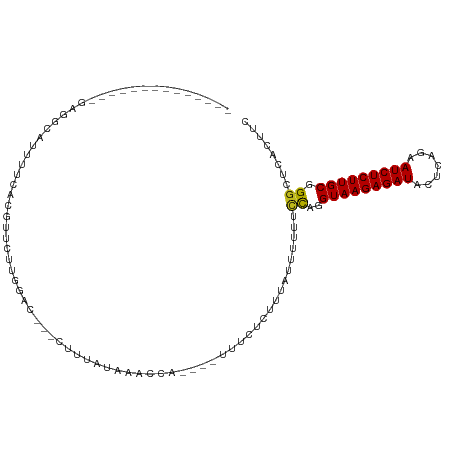

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:20:33 2011