
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 12,115,458 – 12,115,559 |
| Length | 101 |
| Max. P | 0.500000 |

| Location | 12,115,458 – 12,115,559 |
|---|---|
| Length | 101 |
| Sequences | 14 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 89.36 |
| Shannon entropy | 0.25903 |
| G+C content | 0.47171 |
| Mean single sequence MFE | -28.38 |
| Consensus MFE | -21.71 |
| Energy contribution | -21.87 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.34 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3R 12115458 101 - 27905053 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.((((((.(((((.(((((.........))))).)))))..((((...........)))))))))).)).....))))))))) ( -27.70, z-score = -1.69, R) >droSec1.super_12 91218 101 + 2123299 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.((((((.(((((.(((((.........))))).)))))..((((...........)))))))))).)).....))))))))) ( -27.70, z-score = -1.69, R) >droSim1.chr3R 18167270 101 + 27517382 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.((((((.(((((.(((((.........))))).)))))..((((...........)))))))))).)).....))))))))) ( -27.70, z-score = -1.69, R) >droYak2.chr3R 14522645 101 + 28832112 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUUGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.(((((((((((((((((..........))))))).(((...((.....))..))))))))))))).)).....))))))))) ( -27.90, z-score = -1.72, R) >droEre2.scaffold_4770 11639797 101 - 17746568 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGAACACCGGUUGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.((((((((((....(((((((.(((.(((....)))))).)....))))))....)))))))))).)).....))))))))) ( -27.80, z-score = -1.85, R) >droAna3.scaffold_13340 15460780 101 - 23697760 AAACGCAAUUCGCCAAAGUUACCACAGCCAAUUUUUUUGCCAACACGAAAAUUUGGUCCGACCAUAAGGACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......((((((((((((.(((((.((((((((((.((....)).)))))))..((((........))))...))).))))))))......))))))))) ( -31.60, z-score = -2.19, R) >dp4.chr2 8293814 101 + 30794189 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.((((((.(((((.(((((.........))))).)))))..((((...........)))))))))).)).....))))))))) ( -27.70, z-score = -1.69, R) >droPer1.super_3 2095173 101 + 7375914 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.((((((.(((((.(((((.........))))).)))))..((((...........)))))))))).)).....))))))))) ( -27.70, z-score = -1.69, R) >droWil1.scaffold_181130 3311803 101 - 16660200 AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCUGUUGUGGUGCUUACAGAUUGGCGGGU .......(((((((((((.(((((((((.(((((((..........)))))))(((.....)))...........))))))))).)).....))))))))) ( -23.20, z-score = -0.54, R) >droVir3.scaffold_13047 1727936 101 - 19223366 AAACGCAAUUCGCCAAAGUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGCACACCGGUCGUGGUGCUCACAGAUUGGCGGGU .......((((((((((((.(((((.(((((.(((((.........))))).)))))..((((...........))))))))))))......))))))))) ( -28.90, z-score = -1.77, R) >droGri2.scaffold_14906 12544565 101 - 14172833 AAGCGCAAUUCGCCAAAGUUGCCACAGCCAAUUUUUUUGCCAACACGAAAAUUUGGUCCGACCAUAAGGACACCGGUUGUGGUGCUUACAGAUUGGCGGGU .......((((((((((((.((((((((((((((((.((....)).)))))))..((((........))))...))))))))))))......))))))))) ( -34.40, z-score = -2.23, R) >droMoj3.scaffold_6540 18697266 101 + 34148556 AAACGCAAUUCGCCAAAGUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUGAGCACACCGGUUGUGGUGCUUACAGAUUGGCGGGU .......((((((((((((.((((((((((((((((..........)))))))(((.....)))..........))))))))))))......))))))))) ( -28.90, z-score = -1.47, R) >anoGam1.chr2R 49490433 101 - 62725911 AGCCUGAGUUCUCCGAAGUUGCCACAGCCGAUCUUUUUGCCGACGCGGAAGUUGGGGCCCACCAUCAGGACACCGGAGGACGAACUGACGGAAUGGCGUGC .(((...((((((((.....(((....(((((..((((((....))))))))))))))...((....))....))))))))...(.....)...))).... ( -28.00, z-score = 1.60, R) >triCas2.chrUn_2 333224 101 - 1160328 AGGCGUAAUUCGCCGAAGUUUCCACAGCCUAUUUUCUUCCCAACUCGGAAAUUUGGACCCACCAUUAAAACCCCAGAACUGGAGUUGGAGAUAUGGCGUGU .((((.....))))........(((.(((.....(((..(((((((.(...(((((................)))))..).))))))))))...)))))). ( -28.09, z-score = -1.59, R) >consensus AAACGCAAUUCGCCAAAAUUACCACAGCCAAUUUUUUUACCGACACGAAAAUUUGGUCCGACCAUAAGUACACCGGUCGUGGUGCUUACAGAUUGGCGGGU .......(((((((((...((((((.(((((.(((((.........))))).)))))..((((...........))))))))))........))))))))) (-21.71 = -21.87 + 0.17)


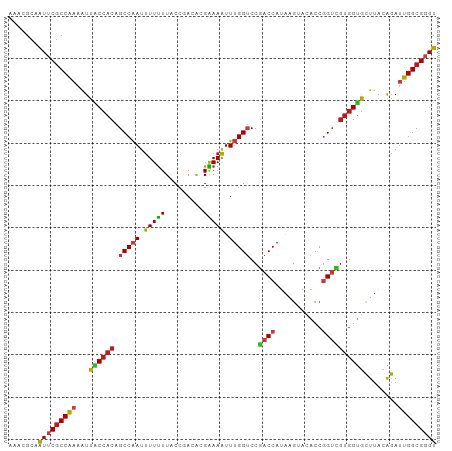
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:19:27 2011