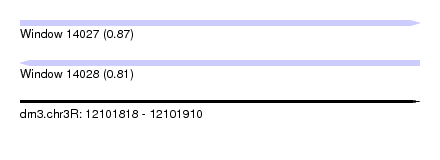

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 12,101,818 – 12,101,910 |
| Length | 92 |
| Max. P | 0.866601 |

| Location | 12,101,818 – 12,101,910 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 88.71 |
| Shannon entropy | 0.18888 |
| G+C content | 0.49007 |
| Mean single sequence MFE | -27.67 |
| Consensus MFE | -20.24 |
| Energy contribution | -21.28 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.866601 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
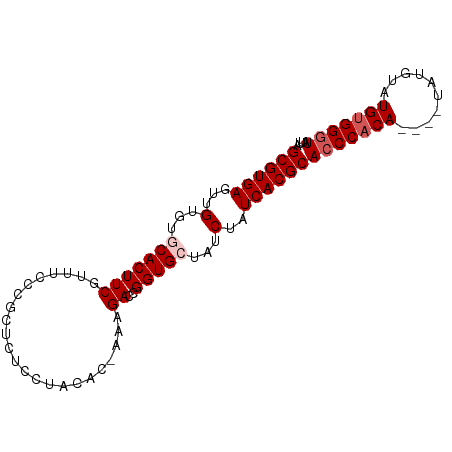
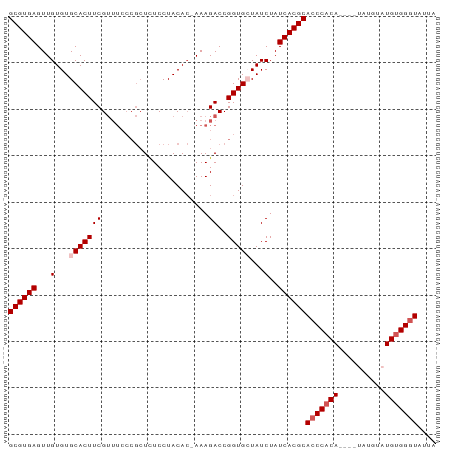
>dm3.chr3R 12101818 92 + 27905053 UAAUACCCACAUACAUACAUAUGUGGGUGCGUGAUAGAUAGCACCGGUCUUUCGUGUAGGAGAGCGGGAAACGAAGUGCACACAACUCACGC ....(((((((((......)))))))))((((((......((((((.(((((......))))).))(....)...)))).......)))))) ( -32.92, z-score = -3.58, R) >droSim1.chr3R 18153387 87 - 27517382 UAAUACCCACAUACAUA----UGUGGGUGCGUGAUAGAUAGCACCGGUCUUU-GUGUAGGAGAGCGGGAAACGAAGUGCACACAACUCACGC ....((((((((....)----)))))))((((((......((((((.(((((-.....))))).))(....)...)))).......)))))) ( -32.22, z-score = -3.66, R) >droSec1.super_12 77326 87 - 2123299 UAAUACCCACAUACAUA----UGUGGGUGCGUGAUAGAUAGCACCGGUCUUU-GUGUAGGAGAGCGGGAAACGAAGUGCACACAACUCACGC ....((((((((....)----)))))))((((((......((((((.(((((-.....))))).))(....)...)))).......)))))) ( -32.22, z-score = -3.66, R) >droYak2.chr3R 14504094 83 - 28832112 UAAUACCCGCAUA--------UGUGGGUGCGUGAUAGAUACCACCGGUCUUU-GUGUAGGAAAGCGGGAAACGAAGUGCACACAACUCACGC ....((((((...--------.))))))((((((.((((.......))))((-((((.(.(...((.....))...).))))))).)))))) ( -25.50, z-score = -1.39, R) >droEre2.scaffold_4770 11626214 75 + 17746568 UAAUAGCCACAUA--------UGUGGGUGCGUGAUAGAUACCACCGGUCUUU-UUGUAGGAAA--------CGAAGUGCACACAACUCACGC ......((((...--------.))))..((((((.((((.......))))..-.....(....--------)..............)))))) ( -15.50, z-score = 0.40, R) >consensus UAAUACCCACAUACAUA____UGUGGGUGCGUGAUAGAUAGCACCGGUCUUU_GUGUAGGAGAGCGGGAAACGAAGUGCACACAACUCACGC ....(((((((..........)))))))((((((.........(((.(((((......))))).)))........((....))...)))))) (-20.24 = -21.28 + 1.04)



| Location | 12,101,818 – 12,101,910 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 88.71 |
| Shannon entropy | 0.18888 |
| G+C content | 0.49007 |
| Mean single sequence MFE | -26.13 |
| Consensus MFE | -20.38 |
| Energy contribution | -21.18 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.813360 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 12101818 92 - 27905053 GCGUGAGUUGUGUGCACUUCGUUUCCCGCUCUCCUACACGAAAGACCGGUGCUAUCUAUCACGCACCCACAUAUGUAUGUAUGUGGGUAUUA ((((((...(((.(((((..((((..((..........))..)))).))))))))...))))))((((((((((....)))))))))).... ( -34.00, z-score = -3.80, R) >droSim1.chr3R 18153387 87 + 27517382 GCGUGAGUUGUGUGCACUUCGUUUCCCGCUCUCCUACAC-AAAGACCGGUGCUAUCUAUCACGCACCCACA----UAUGUAUGUGGGUAUUA ((((((...(((.(((((..((((...............-..)))).))))))))...))))))(((((((----(....)))))))).... ( -28.63, z-score = -2.83, R) >droSec1.super_12 77326 87 + 2123299 GCGUGAGUUGUGUGCACUUCGUUUCCCGCUCUCCUACAC-AAAGACCGGUGCUAUCUAUCACGCACCCACA----UAUGUAUGUGGGUAUUA ((((((...(((.(((((..((((...............-..)))).))))))))...))))))(((((((----(....)))))))).... ( -28.63, z-score = -2.83, R) >droYak2.chr3R 14504094 83 + 28832112 GCGUGAGUUGUGUGCACUUCGUUUCCCGCUUUCCUACAC-AAAGACCGGUGGUAUCUAUCACGCACCCACA--------UAUGCGGGUAUUA ((((((.(((((((.....((.....))......)))))-))..(((...))).....))))))((((.(.--------...).)))).... ( -21.50, z-score = -0.48, R) >droEre2.scaffold_4770 11626214 75 - 17746568 GCGUGAGUUGUGUGCACUUCG--------UUUCCUACAA-AAAGACCGGUGGUAUCUAUCACGCACCCACA--------UAUGUGGCUAUUA ((((((...(.(((((((..(--------(((.......-..)))).))).)))))..))))))..((((.--------...))))...... ( -17.90, z-score = -0.29, R) >consensus GCGUGAGUUGUGUGCACUUCGUUUCCCGCUCUCCUACAC_AAAGACCGGUGCUAUCUAUCACGCACCCACA____UAUGUAUGUGGGUAUUA ((((((...(...(((((((.......................))..)))))...)..))))))(((((((..........))))))).... (-20.38 = -21.18 + 0.80)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:19:24 2011