
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 11,738,796 – 11,738,991 |
| Length | 195 |
| Max. P | 0.815417 |

| Location | 11,738,796 – 11,738,896 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 74.27 |
| Shannon entropy | 0.49300 |
| G+C content | 0.58600 |
| Mean single sequence MFE | -40.42 |
| Consensus MFE | -28.90 |
| Energy contribution | -28.39 |
| Covariance contribution | -0.51 |
| Combinations/Pair | 1.34 |
| Mean z-score | -0.92 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.799487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 11738796 100 + 27905053 ------GCAGUGGAGGAUGUGGCAUCUGCCGAUGUGGCUGCCAUACUAUCAGCAUCCACCGUCAAGUGUUGCACGUGACACGUGGGCUGGCAGGCGGUGGCACUAC ------...((((.....((.(((((....))))).))((((((.((..((((..((((.((((.(((...))).))))..))))))))..))...)))))))))) ( -38.80, z-score = 0.10, R) >droSim1.chr3R 17777654 97 - 27517382 ------GCAGUGGAGGAUGUGGCAUCUGCCGAUGUGGCUGCCAUACUAUCAGCAUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCAC--- ------(((.(((.(((((...))))).))).))).((..((...((..((((..((((.((((((((...))))))))..))))))))..))..))..))..--- ( -41.50, z-score = -1.10, R) >droSec1.super_0 10900666 97 - 21120651 ------GCAGUGGAUGAUGUGGCUUCUGCCGAUGUGGCUGCCAUACUAUCAGCUUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCAC--- ------..(((.((((.((((((....(((.....))).)))))).)))).))).(((((((((((((...))))))))......(((....))))))))...--- ( -39.20, z-score = -0.81, R) >droYak2.chr3R 14133405 97 - 28832112 ---------GCAGUGGAUGUGGCAUCUGCCGAUGUGGCUGCCAUGCUAUCAGCAUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCACUAC ---------...((((...((((....)))).(((.((((((.(((...((((..((((.((((((((...))))))))..))))))))))))))))).))))))) ( -44.40, z-score = -1.68, R) >droEre2.scaffold_4770 11260006 94 + 17746568 ---------GCAGUGGAUGUGGCAUCUGCCGAUGUGGCUGCCACACCAUCAGCAUCCACCGUCAAGUGUGGCACUUGACACGUGGGCUGGCAGGCGGUGGCAC--- ---------((.((((.(((((((.(..(....)..).)))))))))))..))..(((((((((((((...))))))))......(((....))))))))...--- ( -42.60, z-score = -0.80, R) >droAna3.scaffold_13340 16339624 99 + 23697760 ----UUGCAGCGGCUUGUGCCGCUUCUGCUGCUG---CUGCCAUACUUUCAGUAUUCACUGUCAAGUUCUGGGCUUGACAUGUGGGCUGACAAUUGGUGGCACUGC ----..((((((((....)))((....)).))))---)((((((....(((((..((((((((((((.....)))))))).)))))))))......)))))).... ( -43.00, z-score = -3.23, R) >dp4.chr2 20422213 106 + 30794189 GCUGUUGAUGUUGCCGCUGUUGAUGAUUCCGCAGUUGCGGCCAGAGUAUCAGUGUCCACCGUCAAAUGCUGCGUUUGACAUGGGGCCUGACAGGGACUGGCACUGC ((.((......(((.(((((.((....)).))))).)))(((((....((((.((((...((((((((...))))))))...))))))))......))))))).)) ( -37.20, z-score = -0.15, R) >droPer1.super_3 3169422 106 + 7375914 GCUGUUGAUGUUGCCGCUGUUGAUGAGUCCGCAGUUGCGGCCAGAGUAUCAGUGUCCACCGUCAAAUGCUGCGUUUGACAUGGGGCCUGACAGGGACUGGCACUGC ((.((......(((.(((((.((....)).))))).)))(((((....((((.((((...((((((((...))))))))...))))))))......))))))).)) ( -36.70, z-score = 0.27, R) >consensus ______GCAGUGGAGGAUGUGGCAUCUGCCGAUGUGGCUGCCAUACUAUCAGCAUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCACU_C ..................((.(((((....))))).)).(((((....(((((..((((.((((((((...))))))))..)))))))))......)))))..... (-28.90 = -28.39 + -0.51)



| Location | 11,738,796 – 11,738,896 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 74.27 |
| Shannon entropy | 0.49300 |
| G+C content | 0.58600 |
| Mean single sequence MFE | -36.48 |
| Consensus MFE | -19.88 |
| Energy contribution | -19.96 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.815417 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 11738796 100 - 27905053 GUAGUGCCACCGCCUGCCAGCCCACGUGUCACGUGCAACACUUGACGGUGGAUGCUGAUAGUAUGGCAGCCACAUCGGCAGAUGCCACAUCCUCCACUGC------ ((((((.......(((.((((((((.(((((.(((...))).)))))))))..)))).)))..((((((((.....)))...))))).......))))))------ ( -35.30, z-score = -0.64, R) >droSim1.chr3R 17777654 97 + 27517382 ---GUGCCACCGCCUGCCAGCCCACGUGUCAAGUGCAACACUUGACGGUGGAUGCUGAUAGUAUGGCAGCCACAUCGGCAGAUGCCACAUCCUCCACUGC------ ---(((.((.(((((((((((((((.(((((((((...)))))))))))))..))))...))).))).(((.....))).).)).)))............------ ( -36.10, z-score = -1.34, R) >droSec1.super_0 10900666 97 + 21120651 ---GUGCCACCGCCUGCCAGCCCACGUGUCAAGUGCAACACUUGACGGUGGAAGCUGAUAGUAUGGCAGCCACAUCGGCAGAAGCCACAUCAUCCACUGC------ ---(((.(...((((((((((((((.(((((((((...)))))))))))))..))))...))).))).(((.....)))....).)))............------ ( -35.10, z-score = -1.52, R) >droYak2.chr3R 14133405 97 + 28832112 GUAGUGCCACCGCCUGCCAGCCCACGUGUCAAGUGCAACACUUGACGGUGGAUGCUGAUAGCAUGGCAGCCACAUCGGCAGAUGCCACAUCCACUGC--------- ((((((.....(.(((((...((((.(((((((((...)))))))))))))((((.....))))))))).).....(((....))).....))))))--------- ( -40.80, z-score = -2.31, R) >droEre2.scaffold_4770 11260006 94 - 17746568 ---GUGCCACCGCCUGCCAGCCCACGUGUCAAGUGCCACACUUGACGGUGGAUGCUGAUGGUGUGGCAGCCACAUCGGCAGAUGCCACAUCCACUGC--------- ---.((((((.(((...((((((((.(((((((((...)))))))))))))..))))..)))))))))(((.....))).((((...))))......--------- ( -41.20, z-score = -1.57, R) >droAna3.scaffold_13340 16339624 99 - 23697760 GCAGUGCCACCAAUUGUCAGCCCACAUGUCAAGCCCAGAACUUGACAGUGAAUACUGAAAGUAUGGCAG---CAGCAGCAGAAGCGGCACAAGCCGCUGCAA---- ....(((((...(((.((((..(((.(((((((.......))))))))))....)))).))).)))))(---((((.((....))(((....))))))))..---- ( -36.90, z-score = -3.22, R) >dp4.chr2 20422213 106 - 30794189 GCAGUGCCAGUCCCUGUCAGGCCCCAUGUCAAACGCAGCAUUUGACGGUGGACACUGAUACUCUGGCCGCAACUGCGGAAUCAUCAACAGCGGCAACAUCAACAGC ((.(.(((((....((((((...((((((((((.......)))))).))))...))))))..))))))((..(((..((....))..)))..))..........)) ( -33.30, z-score = -0.71, R) >droPer1.super_3 3169422 106 - 7375914 GCAGUGCCAGUCCCUGUCAGGCCCCAUGUCAAACGCAGCAUUUGACGGUGGACACUGAUACUCUGGCCGCAACUGCGGACUCAUCAACAGCGGCAACAUCAACAGC ((.(.(((((....((((((...((((((((((.......)))))).))))...))))))..))))))((..(((..((....))..)))..))..........)) ( -33.10, z-score = -0.47, R) >consensus G_AGUGCCACCGCCUGCCAGCCCACGUGUCAAGUGCAACACUUGACGGUGGAUGCUGAUAGUAUGGCAGCCACAUCGGCAGAAGCCACAUCCGCCACUGC______ ....(((((....(((.(((.((((.(((((((.......)))))))))))...))).)))..))))).........((....))..................... (-19.88 = -19.96 + 0.08)


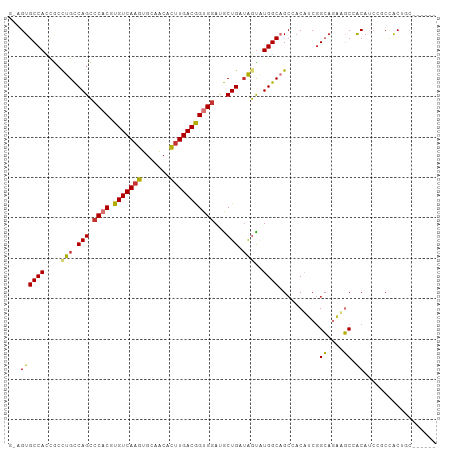
| Location | 11,738,823 – 11,738,930 |
|---|---|
| Length | 107 |
| Sequences | 9 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 76.62 |
| Shannon entropy | 0.46648 |
| G+C content | 0.52087 |
| Mean single sequence MFE | -36.77 |
| Consensus MFE | -19.96 |
| Energy contribution | -18.98 |
| Covariance contribution | -0.98 |
| Combinations/Pair | 1.46 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.753324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 11738823 107 + 27905053 ----GUGGCUGCCAUACUAUCAGCAUCCACCGUCAAGUGUUGCACGUGACACGUGGGCUGGCAGGCGGUGGCACUACU---ACUACUGAGGAUCUGGUUCAAAACUAAAUGCUG ----((((.((((((.((..((((..((((.((((.(((...))).))))..))))))))..))...)))))))))).---.......((.((.((((.....)))).)).)). ( -36.20, z-score = -0.99, R) >droSim1.chr3R 17777681 104 - 27517382 ----GUGGCUGCCAUACUAUCAGCAUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCACUACU---ACU---GAGGAUCUGGUUCAAAACUAAAUGCUG ----((((.((((((.((..((((..((((.((((((((...))))))))..))))))))..))...)))))))))).---...---.((.((.((((.....)))).)).)). ( -40.30, z-score = -2.69, R) >droSec1.super_0 10900693 104 - 21120651 ----GUGGCUGCCAUACUAUCAGCUUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCACUACU---ACU---GAGGAUCUGGUUCAAAACUAAAUGCUG ----((((.((((((.((..((((..((((.((((((((...))))))))..))))))))..))...)))))))))).---...---.((.((.((((.....)))).)).)). ( -40.30, z-score = -2.78, R) >droYak2.chr3R 14133429 110 - 28832112 ----GUGGCUGCCAUGCUAUCAGCAUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCACUACUCGUACUACUGAGGAUCUGGUUCAAAACUAAAUGCUG ----((((.(((((((((.(((((..((((.((((((((...))))))))..)))))))))..))).))))))))))...........((.((.((((.....)))).)).)). ( -39.80, z-score = -1.61, R) >droEre2.scaffold_4770 11260030 104 + 17746568 ----GUGGCUGCCACACCAUCAGCAUCCACCGUCAAGUGUGGCACUUGACACGUGGGCUGGCAGGCGGUGGCACUUCU---ACU---GAGGAUCUGGUUCAAAACUAAAUGCUG ----((((.((((((.((.(((((..((((.((((((((...))))))))..)))))))))..))..))))))...))---)).---.((.((.((((.....)))).)).)). ( -38.10, z-score = -1.22, R) >droAna3.scaffold_13340 16339650 107 + 23697760 ----GCUGCUGCCAUACUUUCAGUAUUCACUGUCAAGUUCUGGGCUUGACAUGUGGGCUGACAAUUGGUGGCACUGCC---ACUGCUGAUGGUCUGGUUCAAAACUAGAUGCUG ----(((((..(((.....(((((..((((((((((((.....)))))))).)))))))))....)))..))).....---..........(((((((.....))))))))).. ( -38.20, z-score = -2.32, R) >dp4.chr2 20422242 111 + 30794189 CGCAGUUGCGGCCAGAGUAUCAGUGUCCACCGUCAAAUGCUGCGUUUGACAUGGGGCCUGACAGGGACUGGCACUGCU---ACUGCUGAUGGUCUGAUUCAAAACAAGAUGCUG .(((((.((.(((((....((((.((((...((((((((...))))))))...))))))))......)))))...)).---))))).....((((...........)))).... ( -38.10, z-score = -0.69, R) >droPer1.super_3 3169451 111 + 7375914 CGCAGUUGCGGCCAGAGUAUCAGUGUCCACCGUCAAAUGCUGCGUUUGACAUGGGGCCUGACAGGGACUGGCACUGCU---ACUGCUGAUGGUCUGAUUCAAAACAAGAUGCUG .(((((.((.(((((....((((.((((...((((((((...))))))))...))))))))......)))))...)).---))))).....((((...........)))).... ( -38.10, z-score = -0.69, R) >droWil1.scaffold_181089 7237874 89 - 12369635 -------------UUGGUCUCAGCAUCCACUGUCAAAUUCUGGCUCUGACAAGGUGG---------GGUGGUACAACU---GCUACUUAUUGAUUGAUUUAAAACUAGAUGCUG -------------.......(((((((((..((((.....))))..)).(((....(---------(((((((....)---))))))).....)))...........))))))) ( -21.80, z-score = -0.93, R) >consensus ____GUGGCUGCCAUACUAUCAGCAUCCACCGUCAAGUGUUGCACUUGACACGUGGGCUGGCAGGCGGUGGCACUACU___ACU_CUGAGGAUCUGGUUCAAAACUAAAUGCUG ....................(((((((((((((((((((...))))))))......(((....))))))))...................((......))........)))))) (-19.96 = -18.98 + -0.98)

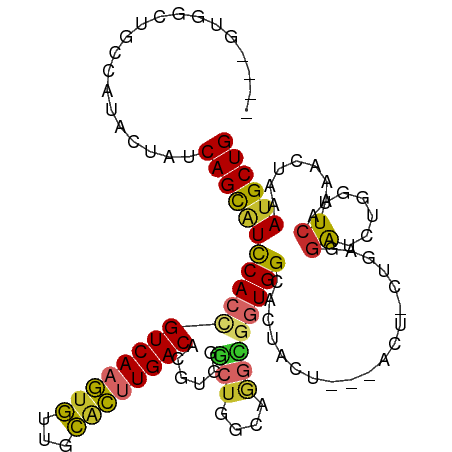

| Location | 11,738,896 – 11,738,991 |
|---|---|
| Length | 95 |
| Sequences | 11 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 66.04 |
| Shannon entropy | 0.66539 |
| G+C content | 0.51727 |
| Mean single sequence MFE | -30.42 |
| Consensus MFE | -14.09 |
| Energy contribution | -14.17 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.57 |
| Mean z-score | -0.88 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.686698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 11738896 95 + 27905053 ---UACUACUGAGGAUCUGGUUCAAAACUAAAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGCAUGGAUGGCAGCAGCCAGGAUGAUGAUGCGGC------------------- ---.........(.(((....(((...((....((((((((..(((..((((((....)))..))).)))...)))))))).))..))).))).)...------------------- ( -27.20, z-score = -0.13, R) >droSim1.chr3R 17777751 95 - 27517382 ---UACUACUGAGGAUCUGGUUCAAAACUAAAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGCAUGGAUGGCAGCAGCCAGGAUGAUGAUGCGGC------------------- ---.........(.(((....(((...((....((((((((..(((..((((((....)))..))).)))...)))))))).))..))).))).)...------------------- ( -27.20, z-score = -0.13, R) >droSec1.super_0 10900763 95 - 21120651 ---UACUACUGAGGAUCUGGUUCAAAACUAAAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGCAUGGAUGGCAGCAGUCAGGAUGAUGAUGCGGC------------------- ---.........(.(((....(((...((....((((((((..(((..((((((....)))..))).)))...)))))))).))..))).))).)...------------------- ( -25.60, z-score = 0.15, R) >droYak2.chr3R 14133502 86 - 28832112 UCGUACUACUGAGGAUCUGGUUCAAAACUAAAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGCAUGGAUGGCAGCAGCCGGCG------------------------------- ........(((......((((.....))))...((((((((..(((..((((((....)))..))).)))...)))))))))))..------------------------------- ( -25.10, z-score = -0.17, R) >droEre2.scaffold_4770 11260100 95 + 17746568 ---UUCUACUGAGGAUCUGGUUCAAAACUAAAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGCAUGGAUGGCAGCAGCCAGUAUGAUGAUCCGGC------------------- ---.........(((((....(((..(((....((((((((..(((..((((((....)))..))).)))...)))))))).))).))).)))))...------------------- ( -32.10, z-score = -1.66, R) >droAna3.scaffold_13340 16339723 97 + 23697760 ---CACUGCUGAUGGUCUGGUUCAAAACUAGAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGUACGGAUGACAGCACUCCGAGUGGCGAUGCGGCAA----------------- ---...(((((.(.(((((...((((((..(((.(((((((.......)))))))))).))))))..))))).))))))......((.((...)).))..----------------- ( -32.70, z-score = -0.84, R) >dp4.chr2 20422319 114 + 30794189 ---UACUGCUGAUGGUCUGAUUCAAAACAAGAUGCUGUUGCAACCAAGGCAACGGCUCCGUUUUGCAUGGAUGACAGCAGCCCGCCGACUGCUGUUGAUGCUGCUGCUCUCGCACUG ---...(((.....((((.((.((((((..((.((((((((.......)))))))))).)))))).))))))(((((((((....((((....))))..))))))).))..)))... ( -38.10, z-score = -1.42, R) >droPer1.super_3 3169528 114 + 7375914 ---UACUGCUGAUGGUCUGAUUCAAAACAAGAUGCUGUUGCAACCAAGGCAACGGCUCCGUUUUGCAUGGAUGACAGCAGCCCGCCGACUGCUGUUGAUGCUGCUGCUCUCGCACUG ---...(((.....((((.((.((((((..((.((((((((.......)))))))))).)))))).))))))(((((((((....((((....))))..))))))).))..)))... ( -38.10, z-score = -1.42, R) >droWil1.scaffold_181089 7237929 83 - 12369635 ---UGCUACUUAUUGAUUGAUUUAAAACUAGAUGCUGUUGCCGCCGAGGCAACGGCUCAGUUUGCUCGAGAUGACAGCAAUGUGUC------------------------------- ---((((..(((((..((((....(((((.((.(((((((((.....))))))))))))))))..))))))))).)))).......------------------------------- ( -27.40, z-score = -2.71, R) >droMoj3.scaffold_6540 18889856 96 - 34148556 ------UGCUGAUGGUCUGAUUCAAAACCAGAUGCUGUUGCAGCCAAGGCAACAGCUCCGUUUUGCAUGGAUGACAGCAGCUGGCCGACUGCUGGACA-GCUG-------------- ------.((((.(.((((.((.((((((..((.((((((((.......)))))))))).)))))).)))))).)))))(((((.(((.....))).))-))).-------------- ( -37.80, z-score = -1.93, R) >anoGam1.chr2R 6564958 82 - 62725911 ---AACUGAUGGUGCCUGUGGUUCAUUUGAUGGACCGCUGCCACCUAAUAACAGUGGAUGGUGUCCAGCGAGGGGGAGAUGCGGC-------------------------------- ---.((((..((((.(.(((((((((...))))))))).).))))......))))(.((..(.(((.....))).)..)).)...-------------------------------- ( -23.30, z-score = 0.57, R) >consensus ___UACUACUGAUGAUCUGGUUCAAAACUAAAUGCUGUUGCAGCCAAGGCAACGGAUCCGUUUUGCAUGGAUGGCAGCAGCCAGCAUGAUGAUGCGGC___________________ ........(((...(((((...((((((......(((((((.......)))))))....))))))..)))))..)))........................................ (-14.09 = -14.17 + 0.08)


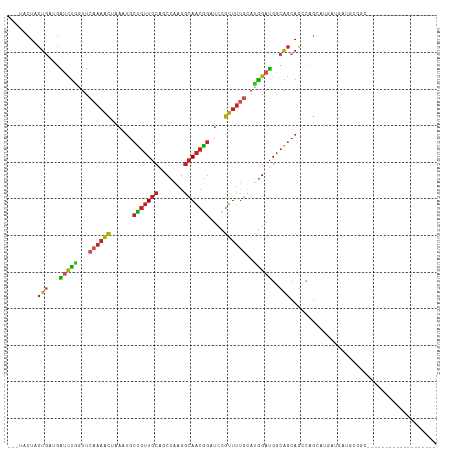
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:18:32 2011