
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 11,341,384 – 11,341,487 |
| Length | 103 |
| Max. P | 0.878182 |

| Location | 11,341,384 – 11,341,487 |
|---|---|
| Length | 103 |
| Sequences | 12 |
| Columns | 122 |
| Reading direction | forward |
| Mean pairwise identity | 68.90 |
| Shannon entropy | 0.59055 |
| G+C content | 0.43203 |
| Mean single sequence MFE | -24.65 |
| Consensus MFE | -12.59 |
| Energy contribution | -12.33 |
| Covariance contribution | -0.26 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.878182 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 11341384 103 + 27905053 -----GGAGUGGAGCU-GUGACUGUGACUG------CAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUGAUGGCGACGACUAUGGGUACAAUGAGG------- -----.......((((-(..(((((....)------).)))..)...((((((((((...........))))))))))))))((((.(.((....)).)...)))).........------- ( -25.20, z-score = -0.51, R) >droSim1.chr3R 17382547 103 - 27517382 -----GGAGUGGAGCU-GUGACUGUGACUG------CAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUGAUGGCGACGACUAUGGGUACAAUGAGG------- -----.......((((-(..(((((....)------).)))..)...((((((((((...........))))))))))))))((((.(.((....)).)...)))).........------- ( -25.20, z-score = -0.51, R) >droSec1.super_0 10516970 103 - 21120651 -----GGAGUGGAGCU-GUGACUGUGACUG------CAAGUUGCAUGAAAUUAUGCUUUAAUCUAUUUAGCAUAAUUUGGCUGCCCUGAUGGCGACGACUAUGGGUACAAUGAGG------- -----.......((((-(..(((((....)------).)))..)...((((((((((...........))))))))))))))((((.(.((....)).)...)))).........------- ( -25.20, z-score = -0.41, R) >droYak2.chr3R 13745587 114 - 28832112 GGAGUGGAGUGGAGCU-GUGACUGUGACUGUGACUGCAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUGAUGGCGACGACUAUGGCUACAAUGAGG------- ...((((.((((((((-.....((..(((((....)).)))..))..((((((((((...........))))))))))))))(((.....))).....))))..)))).......------- ( -30.10, z-score = -1.04, R) >droEre2.scaffold_4770 10868861 103 + 17746568 -----GGAGUGGAGCU-GUGACUGUGACUG------CAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUGAUGGCGACGACUAUGGGUACAAUGAGG------- -----.......((((-(..(((((....)------).)))..)...((((((((((...........))))))))))))))((((.(.((....)).)...)))).........------- ( -25.20, z-score = -0.51, R) >droAna3.scaffold_13340 15955157 87 + 23697760 -----CCAGUCAGACU-GCGACUGUAACUG------CGAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUGCUGGCAAUGAC----------------------- -----((((.(((..(-((((((((....)------).)))))))(.((((((((((...........)))))))))).).....))))))).......----------------------- ( -25.60, z-score = -2.38, R) >dp4.chr2 30333574 79 + 30794189 -----GAAGUGGAACU-GCGACUGCGACUG------UAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUCCUG------------------------------- -----.....(((...-(((.(((((((..------...)))))...((((((((((...........)))))))))))).)))..)))..------------------------------- ( -20.20, z-score = -2.44, R) >droPer1.super_6 5725130 79 + 6141320 -----GAAGUGGAACU-GCGACUGCGACUG------UAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUCCUG------------------------------- -----.....(((...-(((.(((((((..------...)))))...((((((((((...........)))))))))))).)))..)))..------------------------------- ( -20.20, z-score = -2.44, R) >droWil1.scaffold_181089 2569370 100 + 12369635 ---------------GCACAGCUGUAAAUG------UAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGCCAGUCACAA-GACUCCAAAGAGUCAUACAGACACAGUCGGGC ---------------((.(.(((((...((------((.((.((.((((((((((((...........))))))))))..)))).))..-(((((....))))).))))...))))).).)) ( -26.50, z-score = -2.64, R) >droMoj3.scaffold_6540 22229865 93 + 34148556 --------GUGGGGCGAACAGACGCAGCCG------UAAGUUGCAUGAAAUUAUGAUUUAAUCUAAUUAGCAUAAUUUGGCGGCAAUGAUGACGAUGGGCGCUGAGA--------------- --------....((((..((..((((..((------(..(((((...((((((((...((((...)))).)))))))).))))).))).)).)).))..))))....--------------- ( -21.60, z-score = -0.07, R) >droVir3.scaffold_12855 4923131 93 + 10161210 --------GUUGUGUAAGCAGACACAGCUG------UAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCGGCAAUGAUGAUGAUGGGCGCUGAGG--------------- --------(((((((......)))))))..------...(((((...((((((((((...........)))))))))).))))).......................--------------- ( -23.50, z-score = -0.50, R) >droGri2.scaffold_14906 4553625 93 - 14172833 --------AUAGUGCAAACAGACACAGCUG------UAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCGGCAAUGAUGGCGAUGGGCGCUGAGG--------------- --------.((((((...((......((((------(..(((((...((((((((((...........)))))))))).)))))....)))))..)).))))))...--------------- ( -27.30, z-score = -1.90, R) >consensus _____GGAGUGGAGCU_GCGACUGUGACUG______CAAGUUGCAUGAAAUUAUGCUUUAAUCUAAUUAGCAUAAUUUGGCUGCCCUGAUGGCGACGACUAUGGAUA_______________ .......................((((((.........))))))...((((((((((...........))))))))))............................................ (-12.59 = -12.33 + -0.26)


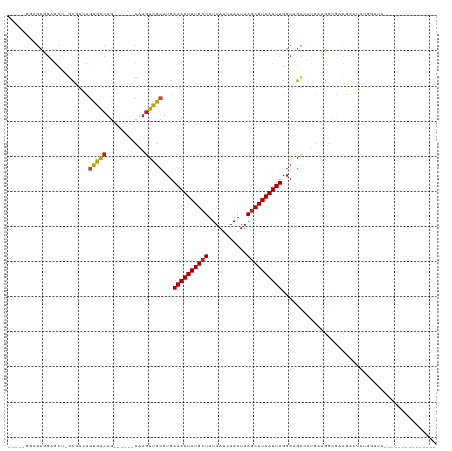
| Location | 11,341,384 – 11,341,487 |
|---|---|
| Length | 103 |
| Sequences | 12 |
| Columns | 122 |
| Reading direction | reverse |
| Mean pairwise identity | 68.90 |
| Shannon entropy | 0.59055 |
| G+C content | 0.43203 |
| Mean single sequence MFE | -18.44 |
| Consensus MFE | -8.62 |
| Energy contribution | -8.79 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.615324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 11341384 103 - 27905053 -------CCUCAUUGUACCCAUAGUCGUCGCCAUCAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUG------CAGUCACAGUCAC-AGCUCCACUCC----- -------....(((((((....(((....(((.....))).((.((((((((((...........))))))))))...)).)))..------..)).)))))...-...........----- ( -17.30, z-score = -0.68, R) >droSim1.chr3R 17382547 103 + 27517382 -------CCUCAUUGUACCCAUAGUCGUCGCCAUCAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUG------CAGUCACAGUCAC-AGCUCCACUCC----- -------....(((((((....(((....(((.....))).((.((((((((((...........))))))))))...)).)))..------..)).)))))...-...........----- ( -17.30, z-score = -0.68, R) >droSec1.super_0 10516970 103 + 21120651 -------CCUCAUUGUACCCAUAGUCGUCGCCAUCAGGGCAGCCAAAUUAUGCUAAAUAGAUUAAAGCAUAAUUUCAUGCAACUUG------CAGUCACAGUCAC-AGCUCCACUCC----- -------....(((((((....(((....(((.....))).((.((((((((((...........))))))))))...)).)))..------..)).)))))...-...........----- ( -17.30, z-score = -0.68, R) >droYak2.chr3R 13745587 114 + 28832112 -------CCUCAUUGUAGCCAUAGUCGUCGCCAUCAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUGCAGUCACAGUCACAGUCAC-AGCUCCACUCCACUCC -------....(((((..(...(((....(((.....))).((.((((((((((...........))))))))))...)).)))....)..))))).........-................ ( -18.20, z-score = -0.21, R) >droEre2.scaffold_4770 10868861 103 - 17746568 -------CCUCAUUGUACCCAUAGUCGUCGCCAUCAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUG------CAGUCACAGUCAC-AGCUCCACUCC----- -------....(((((((....(((....(((.....))).((.((((((((((...........))))))))))...)).)))..------..)).)))))...-...........----- ( -17.30, z-score = -0.68, R) >droAna3.scaffold_13340 15955157 87 - 23697760 -----------------------GUCAUUGCCAGCAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUCG------CAGUUACAGUCGC-AGUCUGACUGG----- -----------------------...(((((.((....(((...((((((((((...........))))))))))..)))..)).)------))))..((((((.-....)))))).----- ( -23.70, z-score = -1.77, R) >dp4.chr2 30333574 79 - 30794189 -------------------------------CAGGAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUA------CAGUCGCAGUCGC-AGUUCCACUUC----- -------------------------------..((((.((.((.((((((((((...........))))))))))..(((.((...------..)).))))).))-..)))).....----- ( -18.70, z-score = -2.43, R) >droPer1.super_6 5725130 79 - 6141320 -------------------------------CAGGAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUA------CAGUCGCAGUCGC-AGUUCCACUUC----- -------------------------------..((((.((.((.((((((((((...........))))))))))..(((.((...------..)).))))).))-..)))).....----- ( -18.70, z-score = -2.43, R) >droWil1.scaffold_181089 2569370 100 - 12369635 GCCCGACUGUGUCUGUAUGACUCUUUGGAGUC-UUGUGACUGGCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUA------CAUUUACAGCUGUGC--------------- ((.((.(((((..((((.(((((....)))))-.........((((((((((((...........)))))))))...)))....))------))..))))).)).))--------------- ( -26.80, z-score = -2.16, R) >droMoj3.scaffold_6540 22229865 93 - 34148556 ---------------UCUCAGCGCCCAUCGUCAUCAUUGCCGCCAAAUUAUGCUAAUUAGAUUAAAUCAUAAUUUCAUGCAACUUA------CGGCUGCGUCUGUUCGCCCCAC-------- ---------------...((((((...((((.....((((....((((((((.((((...))))...))))))))...))))...)------)))..))).)))..........-------- ( -13.70, z-score = 0.03, R) >droVir3.scaffold_12855 4923131 93 - 10161210 ---------------CCUCAGCGCCCAUCAUCAUCAUUGCCGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUA------CAGCUGUGUCUGCUUACACAAC-------- ---------------....((((..(((........((((....((((((((((...........))))))))))...))))....------.....)))..))))........-------- ( -15.43, z-score = -0.83, R) >droGri2.scaffold_14906 4553625 93 + 14172833 ---------------CCUCAGCGCCCAUCGCCAUCAUUGCCGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUA------CAGCUGUGUCUGUUUGCACUAU-------- ---------------.(.((((((.....)).....((((....((((((((((...........))))))))))...))))....------..)))).)..............-------- ( -16.90, z-score = -0.46, R) >consensus _______________UACCCAUAGUCGUCGCCAUCAGGGCAGCCAAAUUAUGCUAAUUAGAUUAAAGCAUAAUUUCAUGCAACUUA______CAGUCACAGUCGC_AGCUCCACUCC_____ .........................................((.((((((((((...........))))))))))...)).......................................... ( -8.62 = -8.79 + 0.17)


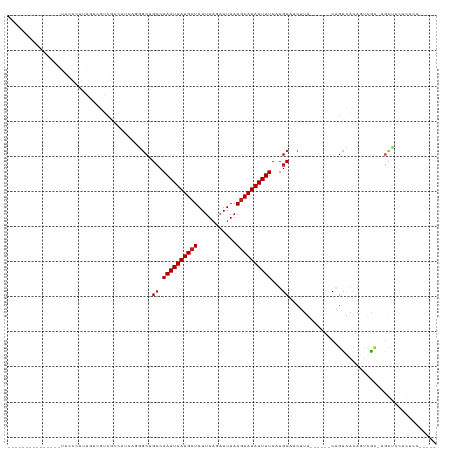
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:17:41 2011