
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 7,513,145 – 7,513,235 |
| Length | 90 |
| Max. P | 0.695658 |

| Location | 7,513,145 – 7,513,235 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 74.11 |
| Shannon entropy | 0.45572 |
| G+C content | 0.49328 |
| Mean single sequence MFE | -26.38 |
| Consensus MFE | -12.31 |
| Energy contribution | -12.77 |
| Covariance contribution | 0.47 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.695658 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 7513145 90 - 23011544 ACAACUCUGUCUUUAGGGUUACCCCCAUGCCCCCAUUCGACAGGCUAAAAUGUCAAUAAAACUAUUGUUGUCCGU---GUCGGAGAGUGACCC----------- ...((((........((((.........))))...((((((((((......))).....(((....))).....)---)))))))))).....----------- ( -19.40, z-score = -0.44, R) >droPer1.super_1 2845171 97 - 10282868 GCAACACUGU-CCUAGGAUUGCCAC----CCGCUUAGCUUUGAGC--AAAUGUCAAUAGUAGUAUUGUUGUCUGUGGCGUCGGAAACGGAGUCCAGAGUGACCC ....((((..-.((.((((((((((----..(((((....)))))--....(.(((((((...))))))).).))))).(((....)))))))))))))).... ( -28.60, z-score = -1.23, R) >dp4.chr4_group3 1377048 97 - 11692001 GCAACUCUGU-CCUAGGAUUGCCAC----CCGCUUAGCUUUGAGC--AAAUGUCAAUAGUAGUAUUGUUGUCUGUGGCGUCGGAAACGGAGUCCAGAGUGACCC ...((((((.-.((......(((((----..(((((....)))))--....(.(((((((...))))))).).))))).(((....)))))..))))))..... ( -30.70, z-score = -1.99, R) >droAna3.scaffold_12916 15448920 84 - 16180835 GCAACUCUGU-UGCAGGGUUGCCCAC---CCACCUCUC--CCGGCUAAAAUGUCAAAAAAAGUUUUGUUGUCCGG---GCCGGGGAGUGACCC----------- (((((((((.-..)))))))))....---.(((.((.(--((((((.......(((((....))))).......)---)))))))))))....----------- ( -32.94, z-score = -2.62, R) >droEre2.scaffold_4929 16433834 83 - 26641161 ACAACUCUGUCUUUAGGGUUGC-------CCCCCAUUCGACAGGCUAAAAUGUCAAUAAAACUAUUGUUGUCCGU---GUCGGGGAGUGACCC----------- ...............((((..(-------(((((((..(((.(((......))).....(((....)))))).))---)..)))).)..))))----------- ( -24.50, z-score = -1.74, R) >droYak2.chr2L 16941361 84 + 22324452 ACAACUCUGUCUUUAGGGUUGCC------CCACCAUGCGACUGGCUAAAAUGUCAAUAAAACUAUUGUUGUCCGU---GUCGGGGAGUGACCC----------- .((((((((....))))))))((------((..((((.(((((((......))))....(((....)))))))))---)..))))........----------- ( -27.70, z-score = -2.58, R) >droSec1.super_3 3029378 84 - 7220098 ACAACUCUGUCUUUAGGGUUGCC------CCACCAUUCGACAGGCUAAAAUGUCAAUAAAACUAUUGUUGUCCGU---GUCGGAGAGUGACCC----------- .((((((((....))))))))..------.(((..((((((((((......))).....(((....))).....)---))))))..)))....----------- ( -23.60, z-score = -2.01, R) >droSim1.chr2L 7309839 84 - 22036055 ACAACUCUGUCUUUAGGGUUGCC------CCACCAUUCGACAGGCUAAAAUGUCAAUAAAACUAUUGUUGUCCGU---GUCGGAGAGUGACCC----------- .((((((((....))))))))..------.(((..((((((((((......))).....(((....))).....)---))))))..)))....----------- ( -23.60, z-score = -2.01, R) >consensus ACAACUCUGUCUUUAGGGUUGCC______CCACCAUUCGACAGGCUAAAAUGUCAAUAAAACUAUUGUUGUCCGU___GUCGGAGAGUGACCC___________ .((((((((....)))))))).....................(((......)))..........(..((.((((......)))).))..).............. (-12.31 = -12.77 + 0.47)

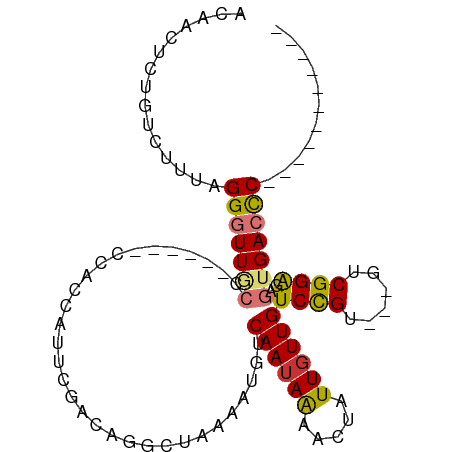

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:23:52 2011