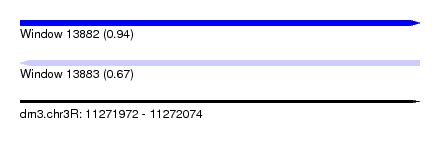

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 11,271,972 – 11,272,074 |
| Length | 102 |
| Max. P | 0.935896 |

| Location | 11,271,972 – 11,272,074 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 64.79 |
| Shannon entropy | 0.69422 |
| G+C content | 0.39441 |
| Mean single sequence MFE | -22.48 |
| Consensus MFE | -10.18 |
| Energy contribution | -10.52 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.935896 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
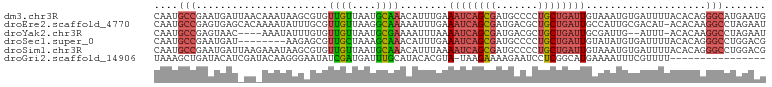
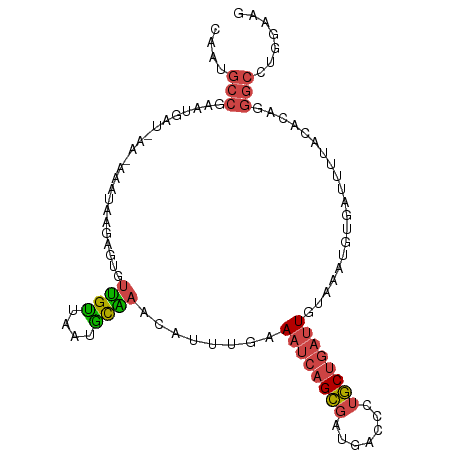
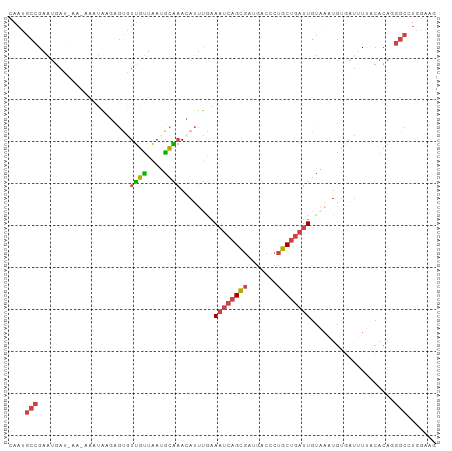
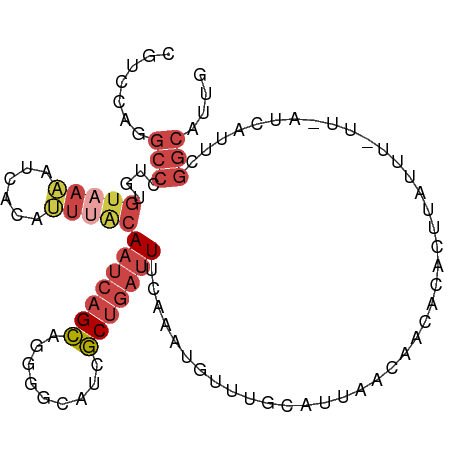
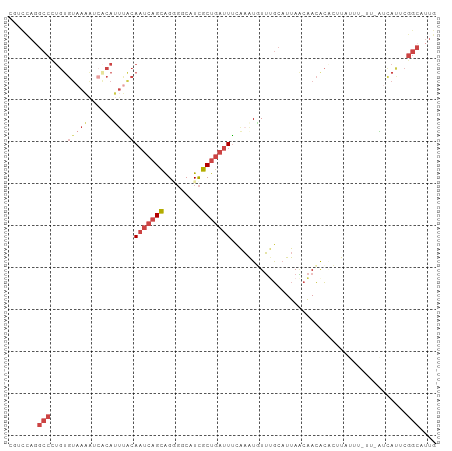
>dm3.chr3R 11271972 102 + 27905053 CAAUGCCGAAUGAUUAACAAAUAAGCGUGUUGUUAAUGCAAACAUUUGAAAUCAGCGAUGCCCCUGCUGAUUGUAAAUGUGAUUUUACACAGGGCAUGAAUG ..(((((...((((((((((.........)))))))).)).(((((((.((((((((.......)))))))).)))))))............)))))..... ( -27.10, z-score = -2.29, R) >droEre2.scaffold_4770 10798843 101 + 17746568 CAAUGCCGAGUGAGCACAAAAUAUUUGCGUUGUUAAGGCAAAAAUUUGAAAUCAGCGAUGACGCUGCUGAUUGCCAUUGCGACAU-ACACAAGGCCUAGAAU ....(((..(((.(((.........)))(((((...((((..........(((((((.......)))))))))))...)))))..-.)))..)))....... ( -27.10, z-score = -1.66, R) >droYak2.chr3R 13673490 95 - 28832112 CAAUGCCGAGUAAC----AAAUAUUUGUGUUGUUAAUGCGAAAAUUUAAAAUCAGCGAUGACGCUGCUGAUUGCGAUUG--AUUU-ACACAAGGCCUAGAAU ....((((((((..----...)))))((((.((((((.((.........((((((((.......)))))))).))))))--))..-))))..)))....... ( -20.80, z-score = -1.19, R) >droSec1.super_0 10443837 94 - 21120651 CAAUGCCGAAUGAU--------AAGAGCGUUGCUAAAGCAAACAUUUGAAAUCAGCGAUGCCCCUGCUGAUUGUAUAUGUGAUUUUACACAGGGCCUGGACG ....(((...((.(--------((((.(.((((....))))((((....((((((((.......))))))))....))))).)))))))...)))....... ( -20.40, z-score = -0.27, R) >droSim1.chr3R 17310726 102 - 27517382 CAAUGCCGAAUGAUUAAGAAAUAAGCGUGUUGUUAAUGCAAACAUUUAAAAUCAGCGAUGCCCCUGCUGAUUGUAAAUGUGAUUUUACACAGGGCCUGGACG ....(((...((..(((((.....((((.......))))..(((((((.((((((((.......)))))))).)))))))..)))))..)).)))....... ( -23.90, z-score = -1.30, R) >droGri2.scaffold_14906 4389120 85 + 14172833 UAAAGCUGAUACAUCGAUACAAGGGAAUAUCGAUGAUUUGCAUACACGUA-UAAGAAAAGAAUCCUCGGCAUGAAAAUUUCGUUUU---------------- ....(((((..((((((((........))))))))...(((......)))-..............)))))................---------------- ( -15.60, z-score = -1.06, R) >consensus CAAUGCCGAAUGAU_AA_AAAUAAGAGUGUUGUUAAUGCAAACAUUUGAAAUCAGCGAUGACCCUGCUGAUUGUAAAUGUGAUUUUACACAGGGCCUGGAAG ....(((......................((((....))))........((((((((.......))))))))....................)))....... (-10.18 = -10.52 + 0.34)
| Location | 11,271,972 – 11,272,074 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 64.79 |
| Shannon entropy | 0.69422 |
| G+C content | 0.39441 |
| Mean single sequence MFE | -21.16 |
| Consensus MFE | -7.03 |
| Energy contribution | -8.53 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.667253 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
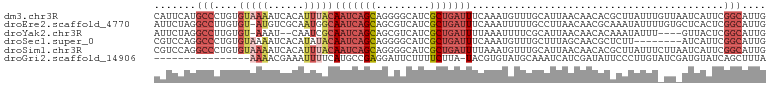
>dm3.chr3R 11271972 102 - 27905053 CAUUCAUGCCCUGUGUAAAAUCACAUUUACAAUCAGCAGGGGCAUCGCUGAUUUCAAAUGUUUGCAUUAACAACACGCUUAUUUGUUAAUCAUUCGGCAUUG .....(((((..((((((....((((((..(((((((.((....)))))))))..))))))))))))((((((.........)))))).......))))).. ( -22.90, z-score = -1.75, R) >droEre2.scaffold_4770 10798843 101 - 17746568 AUUCUAGGCCUUGUGU-AUGUCGCAAUGGCAAUCAGCAGCGUCAUCGCUGAUUUCAAAUUUUUGCCUUAACAACGCAAAUAUUUUGUGCUCACUCGGCAUUG .......((((((((.-....))))).)))......(((((....)))))............((((.......(((((.....))))).......))))... ( -24.54, z-score = -0.99, R) >droYak2.chr3R 13673490 95 + 28832112 AUUCUAGGCCUUGUGU-AAAU--CAAUCGCAAUCAGCAGCGUCAUCGCUGAUUUUAAAUUUUCGCAUUAACAACACAAAUAUUU----GUUACUCGGCAUUG .......(((.((((.-((((--.((((((.....))((((....))))))))....)))).)))).((((((.........))----))))...))).... ( -17.20, z-score = -1.23, R) >droSec1.super_0 10443837 94 + 21120651 CGUCCAGGCCCUGUGUAAAAUCACAUAUACAAUCAGCAGGGGCAUCGCUGAUUUCAAAUGUUUGCUUUAGCAACGCUCUU--------AUCAUUCGGCAUUG .((((..((..((((((........))))))....))..))))...(((((........((((((....)))).))....--------.....))))).... ( -22.63, z-score = -1.29, R) >droSim1.chr3R 17310726 102 + 27517382 CGUCCAGGCCCUGUGUAAAAUCACAUUUACAAUCAGCAGGGGCAUCGCUGAUUUUAAAUGUUUGCAUUAACAACACGCUUAUUUCUUAAUCAUUCGGCAUUG .((((..((....((((((......))))))....))..))))...(((((..((((..((..((...........))..))...))))....))))).... ( -21.60, z-score = -1.33, R) >droGri2.scaffold_14906 4389120 85 - 14172833 ----------------AAAACGAAAUUUUCAUGCCGAGGAUUCUUUUCUUA-UACGUGUAUGCAAAUCAUCGAUAUUCCCUUGUAUCGAUGUAUCAGCUUUA ----------------.....((.((((.(((((((((((......)))).-..)).))))).))))(((((((((......)))))))))..))....... ( -18.10, z-score = -2.27, R) >consensus CGUCCAGGCCCUGUGUAAAAUCACAUUUACAAUCAGCAGGGGCAUCGCUGAUUUCAAAUGUUUGCAUUAACAACACACUUAUUU_UU_AUCAUUCGGCAUUG .......(((....(((((......)))))(((((((.........)))))))..........................................))).... ( -7.03 = -8.53 + 1.50)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:17:25 2011